Page History
Policies
LCLS users are responsible for complying with the data management and curation policies of their home institutions and funding agents and authorities. To enhance the scientific productivity of the LCLS user community, LCLS supplies on-site disk, tape and compute resources for prompt analysis of LCLS data, and software to access those resources consistent with the published data retention policy. Compute resources are preferentially allocated to recent and running experiments.
Data Management
LCLS provides space for all your experiment's data at no cost for you. This includes the raw data from the detectors as well as the data derived from your analysis. Your raw data are available as XTC files or, on demand, as HDF5 files. The tools for managing files are described here.
Getting an Account
You will need a valid SLAC UNIX account in order to use the LCLS computing system. The instructions for getting a SLAC UNIX account are here:
http://www-ssrl.slac.stanford.edu/lcls/users/logistics.html#compaccts
Your UNIX account must be enabled in the LCLS system in order to have access to data and elog. This happens automatically if your account is created with XU as its primary group. If your primary UNIX group is not XU, make a request of enabling your account in the LCLS system by sending an email to:
If you forgot your password or if your account has been disabled send an email to:
account-services@slac.stanford.edu
Getting Access to the System
You can get into the LCLS photon computing system in the NEH by ssh'ing to one of these nodes:
psexport.slac.stanford.edu
psimport.slac.stanford.edu
From these nodes you can move data files in and out of the system and you can connect to the bastion hosts:
pslogin
psdev
Note that, from within SLAC, you can directly connect to the bastion hosts without going through psimport/psexport.
The SLAC wireless visitor network is not considered part of SLAC so you'll need to go through psexport/psimport when using your laptop on-site.
From the bastion hosts you can then reach the analysis nodes (see below).
Each control room has a number of nodes for local login. These nodes have access to the Internet and are named psusr<id>.
The controls and DAQ nodes used for operating an instrument work in kiosk mode so you don't need a personal account to run an experiment from the control room. Remote access to these nodes is not allowed for normal users.
Running the Analysis
The analysis framework is documented in the Data Analysis page for the LCLS-I/HXR systems and psana for the LCLS-II (SXR&UED) systems. This section describes the nodes which are resources available for running the analysis.
Instrument Dedicated Nodes
These nodes are reserved for the users who are currently running an experiment. Each instrument has three dedicated interactive compute systems:
AMO | | SXR | | XPP | |
XCS | | CXI | | MEC | |
The general specifications for these nodes are:
psana<instr>01: 8 cores, Xeon E5520, 24GB, 500GB disk, 1Gb/s, dedicated Matlab licensepsana<instr>02: 8 cores, Xeon E5520, 24GB, 500GB disk, 1Gb/spsana<instr>03: 8 cores, Opteron 2384, 8GB, diskless, 10Gb/s
Interactive Farm
In order to get access to the interactive farm, connect to the address psana. A load-balancing mechanism will connect you to the least loaded of the nodes in the farm:
| No Format |
|---|
ssh psana
|
This farm is currently made of six servers with the following general specifications:
- 8-cores, Opteron 2384, 8GB, diskless, 10Gb/s
Batch Farm
Login first to psdev or pslogin (from SLAC) or psimport or psexport (from anywhere). From there you can submit a job with the following command:
| No Format |
|---|
bsub -q lclsq -o <output file name> <job_script_command>
|
For example:
| No Format |
|---|
bsub -q lclsq -o ~/output/job.out my_program
|
This will submit a job (my_program) to the queue lclsq and write it's output to a file named ~/output/job.out.
You may check on the status of your jobs using the bjobs command.
You can find more LCLS specific information about LSF in this PDF file. For a more detailed description and more LSF commands, please see:
http://www.slac.stanford.edu/comp/unix/unix-hpc.html
The batch farm is made of forty eight servers with the following general specifications:
- 24 cores, Xeon X5675, 24GB memory, 500GB disk, 1Gb/s
Running OpenMPI Jobs in the PCDS Batch Farm
The RedHat supplied OpenMPI packages are installed on psdev, pslogin, psimport, psexport and all of the psana batch servers.
The system default has been set to the current version as supplied by RedHat.
| No Format |
|---|
$ mpi-selector --query
default:openmpi-1.4-gcc-x86_64
level:system
|
Your environment should be set up to use this version (unless you have used RedHat's mpi-selector script, or your login scripts, to override the default). You can check to see if your PATH is correct by issuing the command which mpirun . Currently, this should return: /usr/lib64/openmpi/1.4-gcc/bin/mpirun Future updates to the MPI version may change the exact details of this path.
In addition, your LD_LIBRARY_PATH should include: /usr/lib64/openmpi/1.4-gcc/lib (or something similar).
For notes on compiling examples; please see:
http://www.slac.stanford.edu/comp/unix/farm/mpi.html
The following are examples of how to submit OpenMPI jobs to the PCDS batch queues:
| No Format |
|---|
bsub -q lclsq -a mympi -n 32 -o ~/output/%J.out ~/bin/hello
|
Will submit an OpenMPI job (-a mympi) requesting 32 processors (-n 32) to the lclsq batch queue (-q lclsq).
| No Format |
|---|
bsub -q psfehq -a mympi -n 16 -R "span[ptile=1]" -o ~/output/%J.out ~/bin/hello
|
| Wiki Markup |
|---|
Will submit an OpenMPI job (-a mympi) requesting 18 processors (-n 18) spanned as one processor per host (-R "span\[ptile=1\]") to the psfehq batch queue (-q psfehq). |
Data Storage
LCLS provides space for all your experiment's data at no cost for you. This includes the measurements as well as derived data from your analysis software.
Your data are available as XTC files or, on demand, as HDF5 files.
Short-term Storage
All your data is available on disk for one year after data taking. The path name is /reg/d/psdm. The data files are currently stored in a Lustre file system. Each experiment is allocated three directories: xtc, scratch and hdf5. The xtc directory contains the raw data from the DAQ system, hdf5 directory is for data files in HDF5 format. Contents of xtc and hdf5 directories are archived to tape. The scratch directory is not backed up. Please write the output of your analysis to the scratch area and not in your NFS space. Keep your analysis code under your NFS home or under your NFS group space (if you have one). Your NFS space is backep up.
Long-term Storage
After one year, your data files are removed from disk. The XTC and HDF5 files remain stored on tape for up to 10 years. LCLS may restore your data from tape back to disk for you to access. Restoring the data to disk more than once will require the approval of the LCLS management.
Data Export
There is a web interface to the experimental data accessible via
https://pswww.slac.stanford.edu/apps/explorer
The web interface also allows you to generate file lists that can be fed into bbcp to export your data from SLAC to your home institution. You can use psexport or psimport for copying your data.
See the Data Exportation page for more information.
Printing
The following printers are available in the NEH building from all the UNIX nodes:
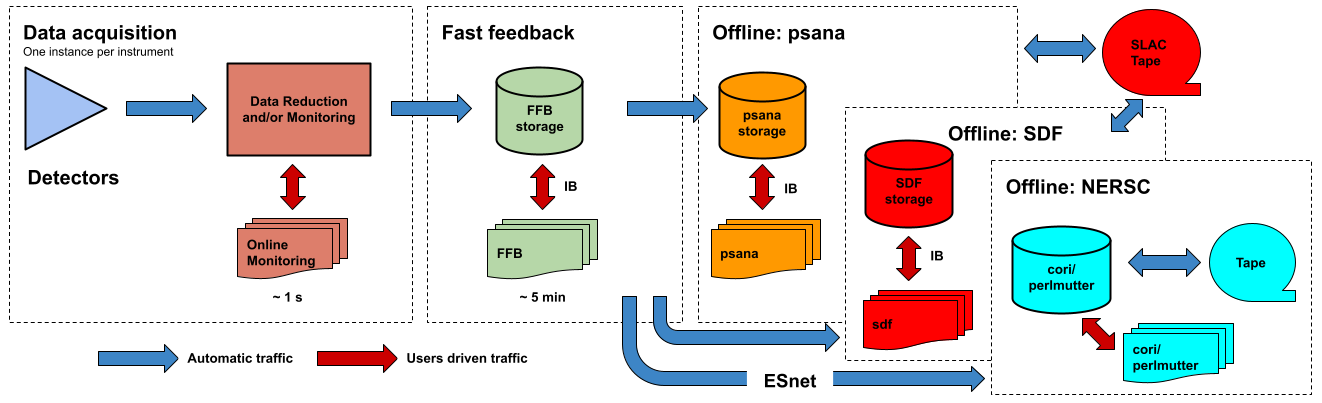
The following figure shows a logic diagram of the LCLS data flow and indicates the different stages where data analysis can be performed in LCLS:
- Data acquisition - The online monitoring nodes get detector data over the network and place it in shared memory on each node. There is a set of monitoring nodes allocated for each instrument. The detector data are received over the network by snooping the multicast traffic between the DAQ readout nodes and the data cache layer. Analysis performed at this stage provides < 1 s feedback capabilities. The methods for doing (quasi) real time analysis are described in the Prompt Analysis page. Users should be aware of the different possibilities and choose the approach that works best for their experiment.
- Fast feedback - The processing nodes in the FFB system read the detector data from a dedicated, flash-based file system. It is possible to read the data as they are written to the FFB storage layer by the DAQ without waiting for the run to end. Analysis performed at this stage provides < 5 min feedback capabilities. The resources reserved for this stage are described in the Fast Feedback System page.
- Offline - The offline nodes read the detector data from disk. These systems include both interactive nodes and batch queues and are the main resource for doing data analysis. We currently support sending the data to three offline systems: psana, S3DF and NERSC. The psana system is the default offline system and your data will end up in psana unless you arrange a different destination with your experiment POC. The psana system is also relatively old and it will be retired when more storage becomes available in the S3DF system. Please consider running at NERSC if you expect to have intensive computing requirements (> 1 PFLOPS).
...
Info
...
Location
...
Device URI
...
Dell 3130
...
AMO Control Room
...
lpd://dellcolor-neh-amo1/lp
...
Dell 3130
...
AMO Control Room
...
lpd://dellcolor-neh-amo2/lp
...
Dell 3130
...
SXR Control Room
...
lpd://dellcolor-neh-sxr1/lp
...
Dell 3130
...
SXR Control Room
...
lpd://dellcolor-neh-sxr2/lp
...
Dell 3130
...
XPP Control Room
...
lpd://dellcolor-neh-xpp1/lp
...
Dell 3130
...
XPP Control Room
...
lpd://dellcolor-neh-xpp2/lp
...
HP Color LaserJet CP3525
...
Bldg 950 corridor ground floor
...
ipp://hpcolor-neh-corridor/ipp/
...
Xerox WorkCentre 5675
...
Bldg 950 Rm 218, Jason Alpers
...
ipp://hpcolor-neh-laser/ipp/
...
HP Color LaserJet 4700
...
Bldg 950 Rm 204, Ray Rodriguez
...
ipp://hpcolor-neh-ray/ipp/
...
HP LaserJet 4350
...
Bldg 950 Rm 203
...