The S3DF (SLAC Shared Science Data Facility) is a new SLAC-wide computing facility. The S3DF will replace the PCDS current computer and storage resources used for the LCLS experiment data processing.
- The S3DF (/sdf/) and PCDS (/cds/) should be considered separate sites as currently storage and home folders are not accessible between each other.
- The S3DF has its own users home directories, and the PCDS home directories (/cds/home/) are not accessible in the S3DF.
- The data directories in the S3DF (/sdf/data/lcls/ds) are not accessible in PCDS and vice versa the PCDS data folders (/cds/data/psdm or /reg/d/psdm) are not accessible in the S3DF.
Accessing the S3DF
To access to the S3DF systems connect to the s3dflogin pool and login with you unix account:
% ssh s3dflogin.slac.stanford.edu
Within the S3DF the nodes are named sdf*. The login nodes don't have the data folders mounted and no Slurm tools. For data transfers, data transfer nodes (s3dfdtn.slac.stanford.edu) are provided.
Interactive cluster
An interactive cluster of nodes is available using the psana name. The interactive node provide access to the LCLS data and allows submitting Slurm jobs.
sdfloginNNN % ssh psana # interactive nodes, same name as in PCDS but different nodes
Data and Software
The data-systems group folder, /sdf/group/lcls/ds, contains software packages and tools with several sub-folders:
| /sdf/group/lcls/ds/ana | psana1/psana2 release, detector calibration,.. |
| /sdf/group/lcls/ds/tools | smalldata-tools, cctbx, crystfel, om, .... |
| /sdf/group/lcls/ds/dm | data-management releases and tools. |
LCLS experimental data
The experiments analysis storage and fast-feed-back storage is accessible on the interactive and batch nodes.
| analysis-storage | /sdf/data/lcls/ds/<instr>/<expt>/<expt-folders> |
| FFB-storage | /sdf/data/lcls/drpsrcf/ffb/<instr>/<expt>/<expt-folders> |
The psana1/psana2 "psconda.sh" scripts shown below set this location for you using this first environment variable (but can also be done with an extra "dir=" DataSource argument). If you are doing realtime FFB analysis you will have to override that with the second value shown here:
export SIT_PSDM_DATA=/sdf/data/lcls/ds/ # offline analysis export SIT_PSDM_DATA=/sdf/data/lcls/drpsrcf/ffb # FFB analysis
S3DF Documentation
S3DF-maintained facility documentation can be found here.
Using psana in S3DF
To access data and submit to Slurm use the interactive cluster (ssh psana).
psana1
% source /sdf/group/lcls/ds/ana/sw/conda1/manage/bin/psconda.sh [-py2]
The command above activates by default the most recent python 3 version of psana. New psana versions do not support python 2 anymore, but it still possible to activate the last available python 2 environment (ana-4.0.45) using the -py2 option.
psana2
% source /sdf/group/lcls/ds/ana/sw/conda2/manage/bin/psconda.sh
Batch processing
The S3DF batch compute link describes the Slurm batch processing. Here we will give a short summary relevant for LCLS users.
a partition and slurm account should be specified when submitting jobs. The slurm account would be lcls:<experiment-name> e.g. lcls:xpp123456 . The account is used for keeping track of resource usage per experiment
% sbatch -p milano --account lcls:xpp1234 ........
- In the S3DF by default memory is limited to 4GB/core. Usually that is not an issue as processing jobs use many core (e.g. a job with 64 cores would request 256GB memory)
- the memory limit will be enforced and your job will fail with an OUT_OF_MEMORY status
- memory can be increased using the --mem sbatch option
- Default total run time is 1 day, the --time option allows to increase/decrease it.
Jupyter
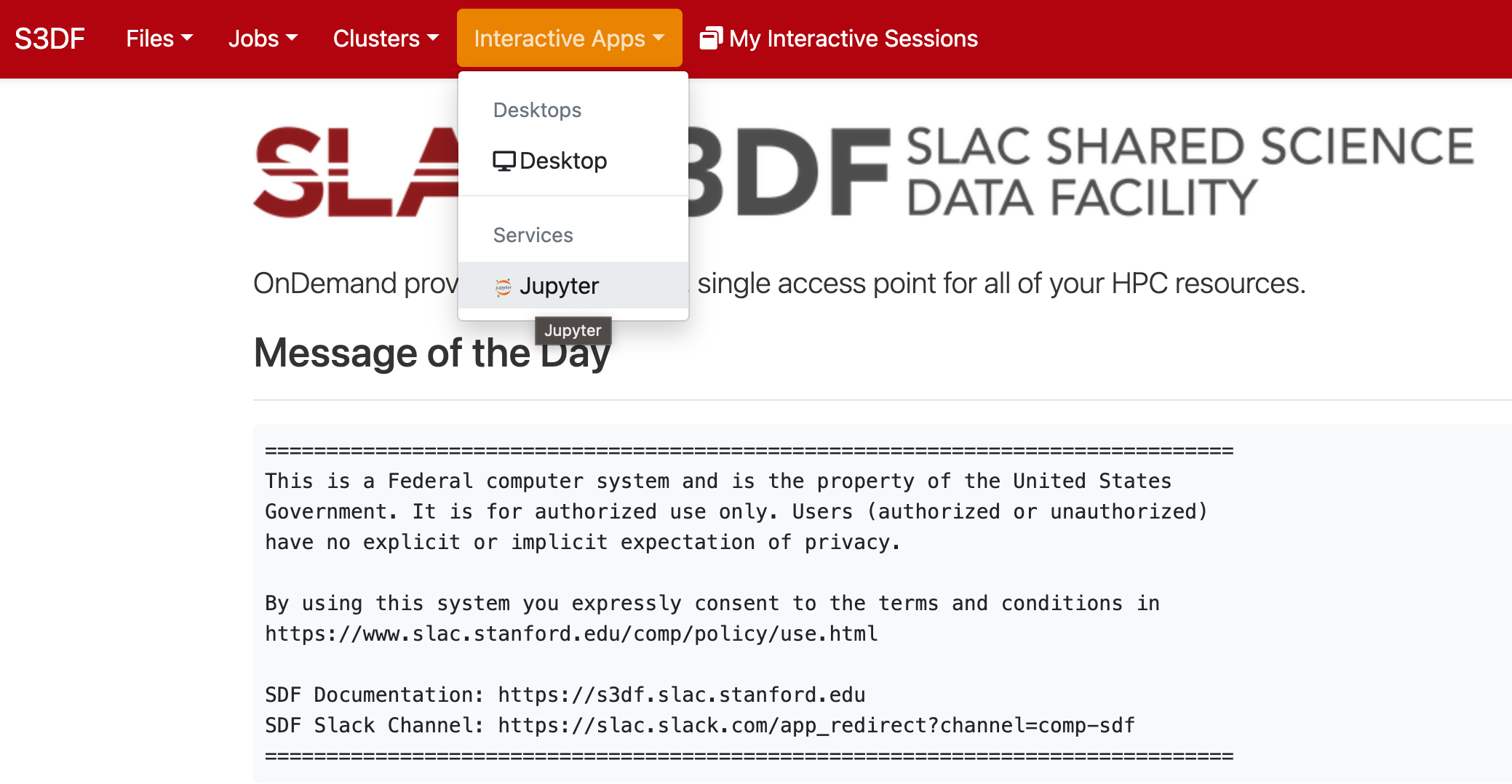
Jupyter is provided by the onDemand service (We are not planning to run a standalone jupyterhub as is done at PCDS. For more information check the S3DF interactive compute docs).
onDemand requires a valid ssh-key (~/.ssh/id_rsa) which are currently not auto generated when an account is enabled in the S3DF. If you don't have a key run the following command on a login or interactive node: /sdf/group/lcls/ds/dm/bin/generate-keys.sh
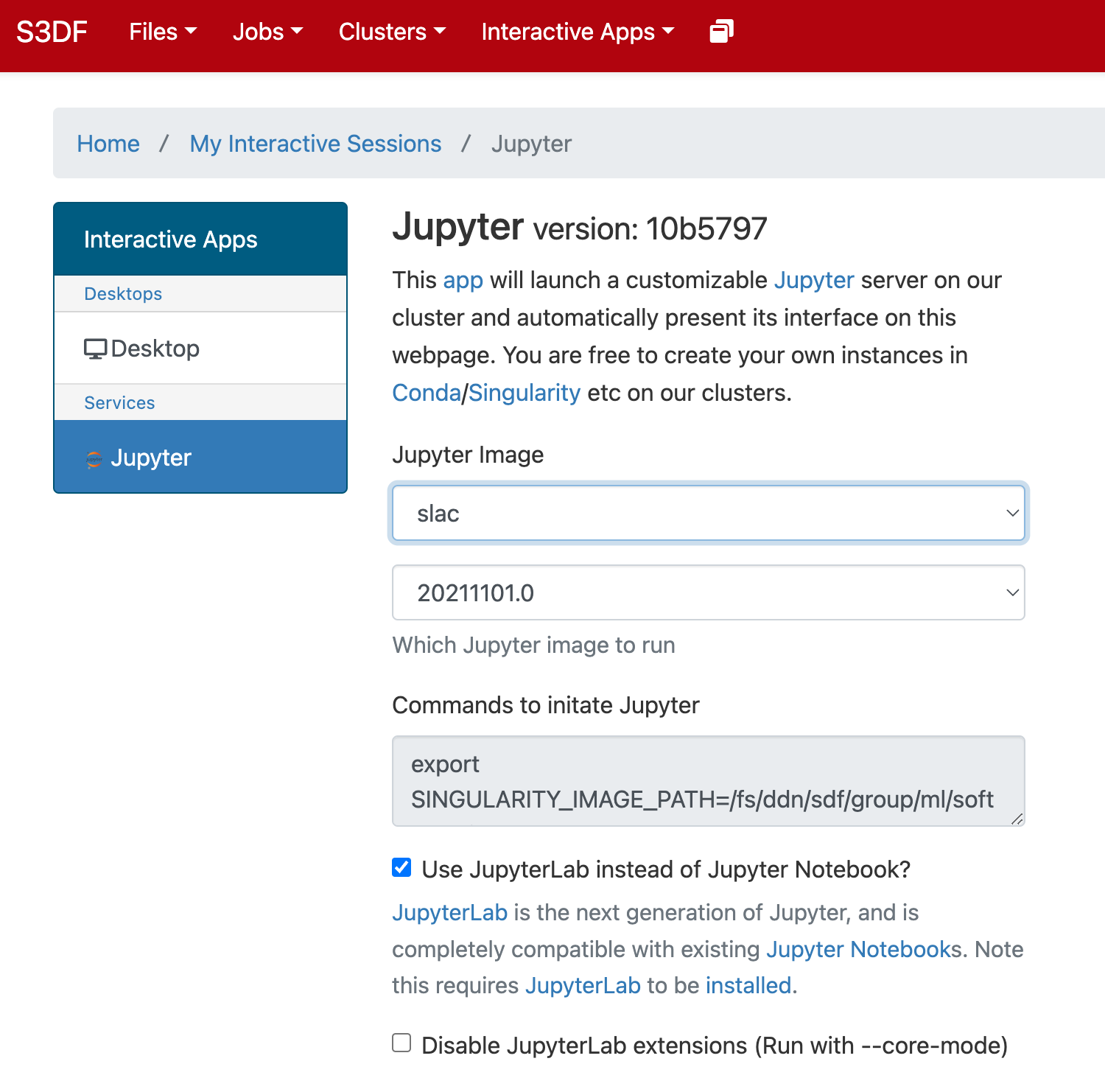
In the "Interactive Apps" select Jupyter, this opens the following form
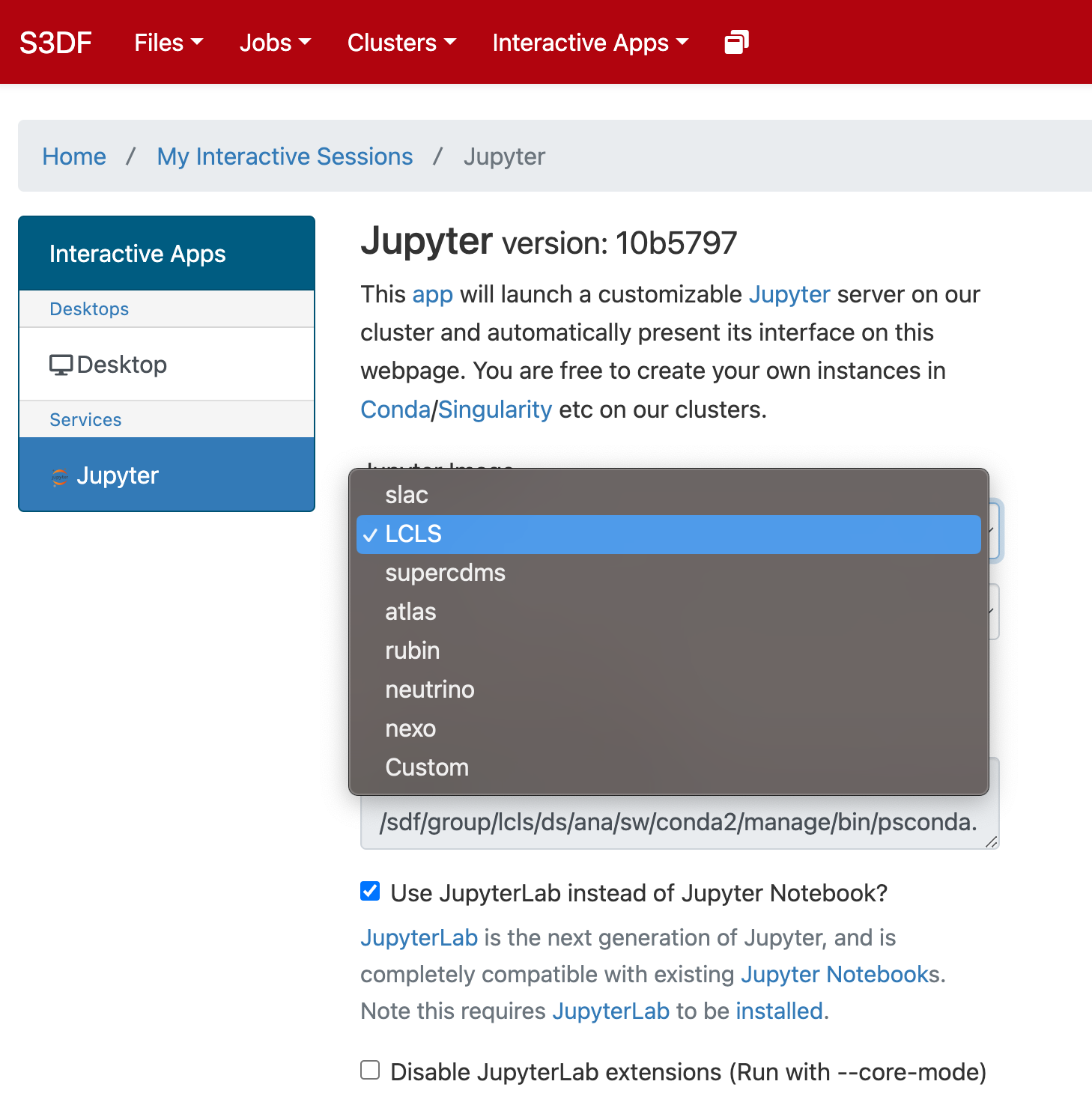
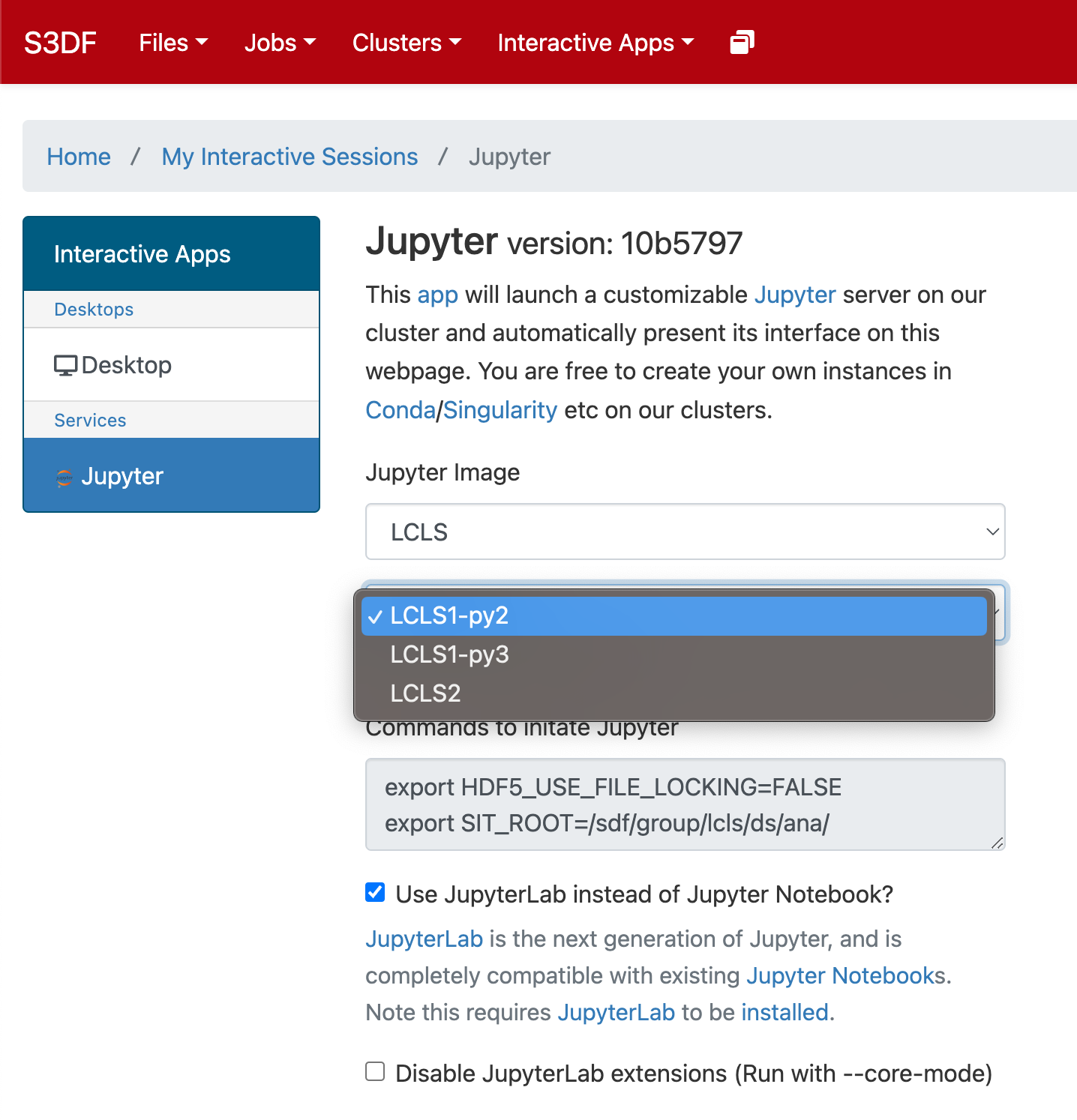
Select the Jupyter image "LCLS" and in the second selection choose "LCLS1-py2", "LCLS1-py3", or "LCLS2".
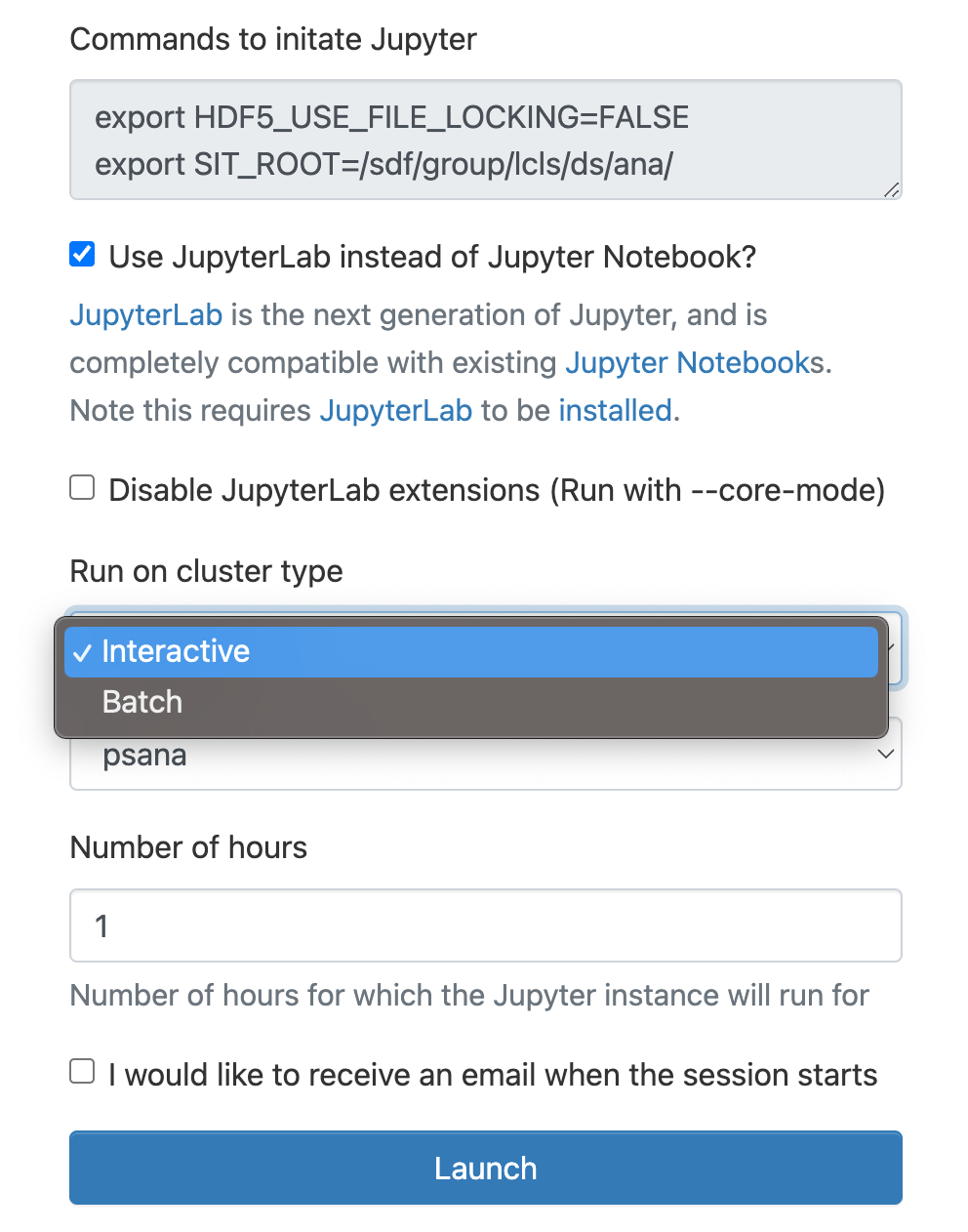
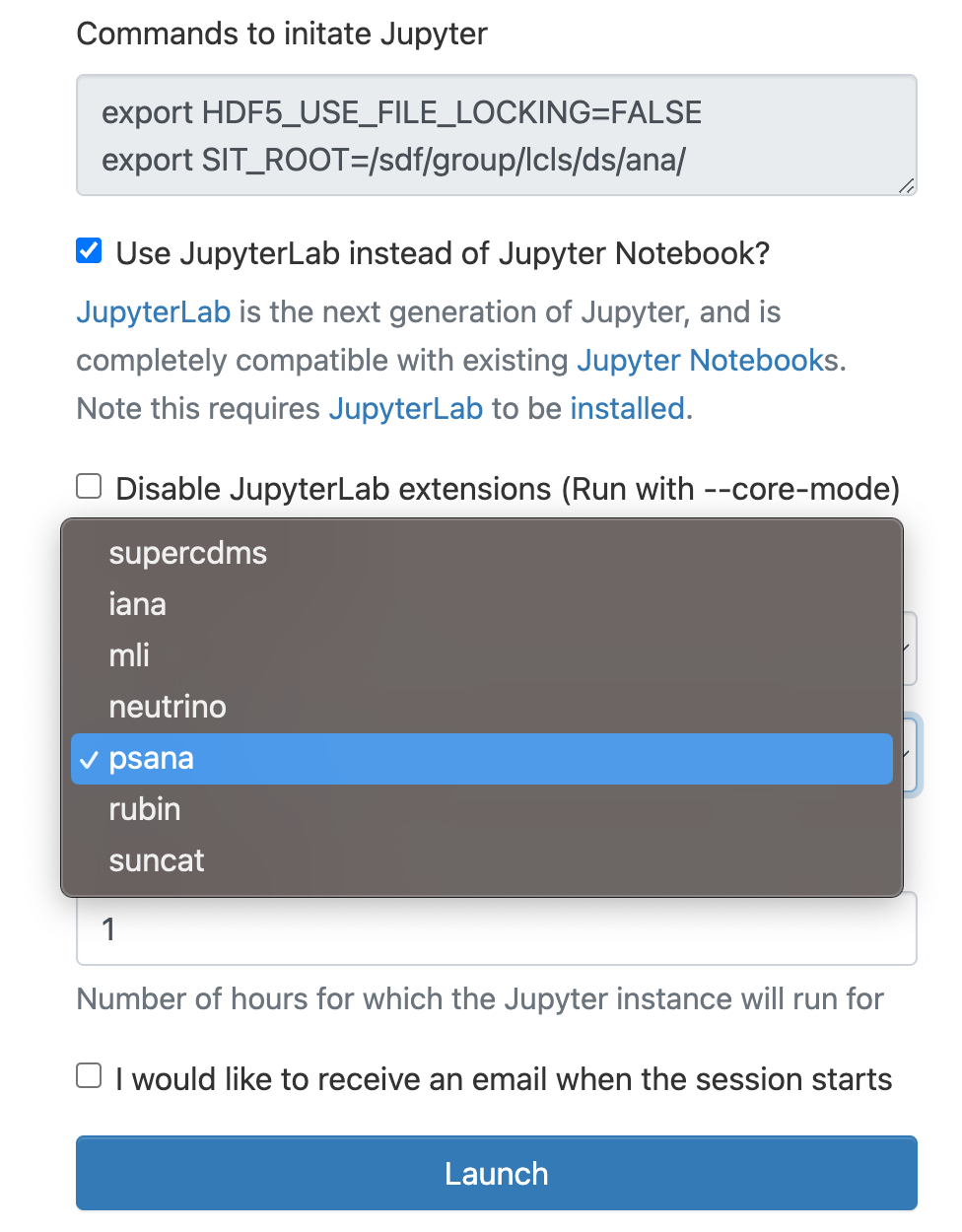
As a cluster type select "Interactive" or "Batch", and then select "psana"(one should always select psana when running LCLS).
Select then how many hours to run and if you prefer Jupyter lab or notebook.
Then click on the "Launch" button:
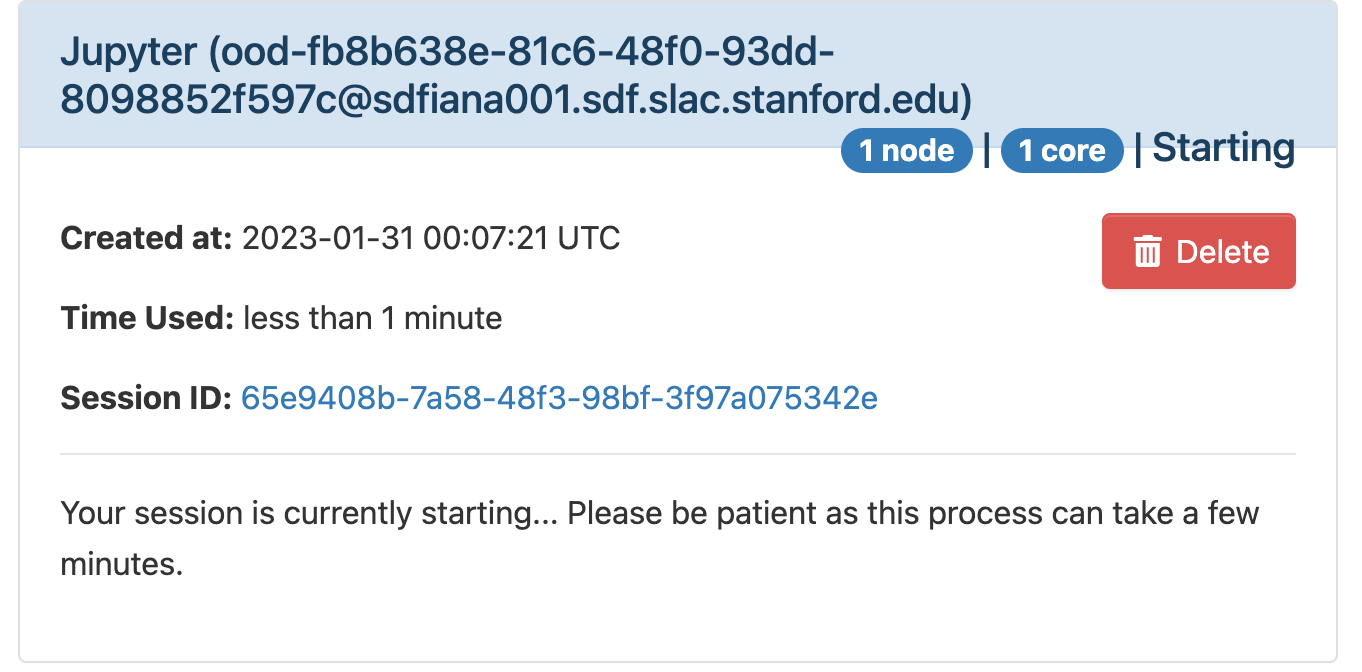
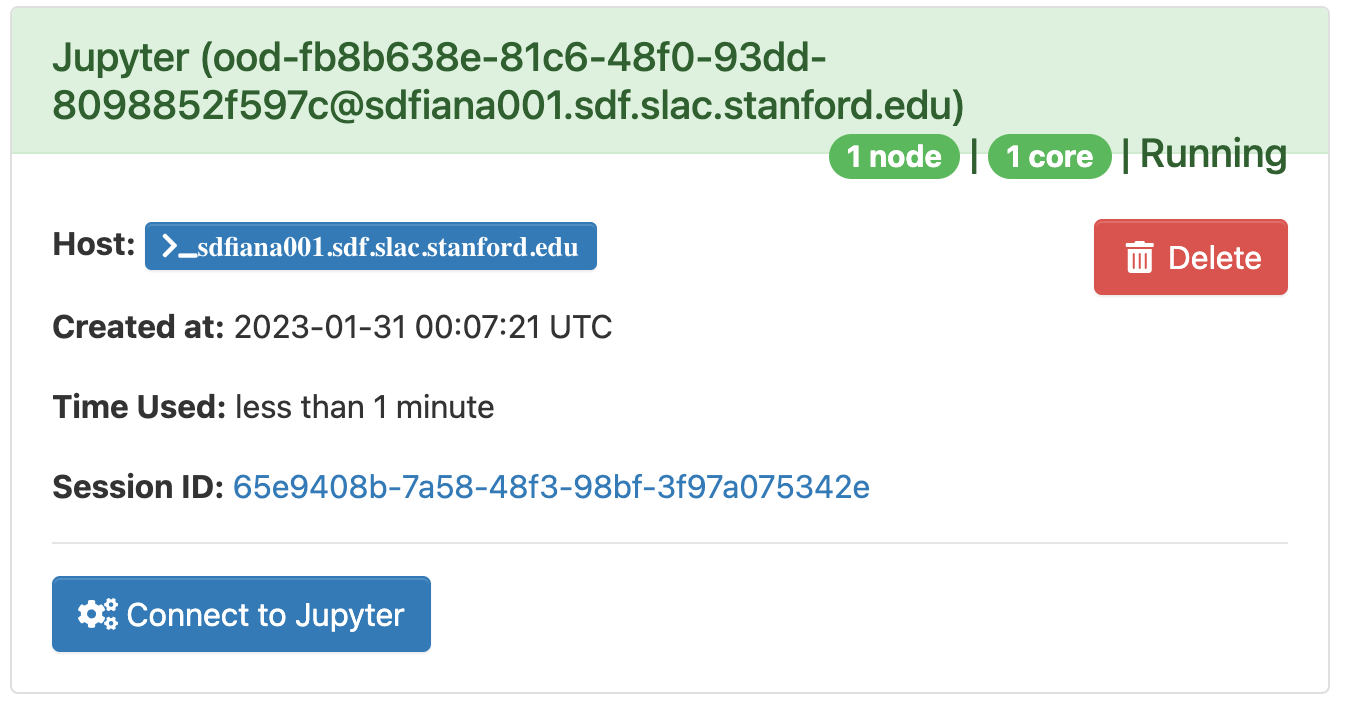
The page will at first show a "Starting" status, after some time it will switch to "Running" and the "Connect to Jupyter" button will be available. Click on the "Connect to Jupyter" button to start working on Jupyter. The Session ID link instead allows to open the directory where the Jupyter is run and access to the files, included the logs.