...
...
...
...
...
...
...
...
- 2011)
- Prior distributions on fit parameters (4 February 2011)
- User defined error and contour level in Minos (3 May 2010)
- Composite2 (11 July 2010)
- RadialProfile model (5 July 2010)
- gtsrcprob tool: Presentation to 14 May 2010 WAM (See also LK-57@JIRA.)
- Upper limits using the (semi-)Bayesian method of Helene (21 September 2009)
- Interface to MINOS errors from Minuit or NewMinuit (29 June 2009)
- EBL Attenuation models (20 June 2009)
- SummedLikelihood (11 June 2009)
- SmoothBrokenPowerLaw (11 June 2009)
- Flux, energy flux, and upper limit calculations for diffuse sources (22 Feb 2009)
- Diffuse response calculations and FITS image templates (22 Feb 2009)
- Analysis object creation using the hoops/ape interface
- Photon and Energy Flux Calculations
Calculating SEDs directly from pyLikelihood (8 November 2011)
SEDs can be computed directly from pyLikelihod using a new SED class.
This code has the advantage that it does not create any temporary files and does not require multiple gtlike jobs or xml files. The only additional dependency of the code is scipy.
Usage example:
| Code Block |
|---|
|#Calculating SEDs directly from pyLikelihood (8 November 2011)] * [Prior distributions on fit parameters (4 February 2011)|#Prior distributions on fit parameters (4 February 2011)] * [User defined error and contour level in Minos (3 May 2010)|#User defined error and contour level in Minos (3 May 2010)] * [Composite2 (11 July 2010)|#Composite2 (11 July 2010)] * [RadialProfile model (5 July 2010)|#RadialProfile model (5 July 2010)] * gtsrcprob tool: [Presentation|https://confluence.slac.stanford.edu/download/attachments/75006028/ST_dev_WAM_13May10.pdf?version=1&modificationDate=1273820184000] to [14 May 2010 WAM|SCIGRPS:Weekly Analysis Meeting - 2010] (See also [LK-57@JIRA].) * [Upper limits using the (semi-)Bayesian method of Helene (21 September 2009)|#Upper limits using the (semi-)Bayesian method of Helene (21 September 2009)] * [Interface to MINOS errors from Minuit or NewMinuit (29 June 2009)|#Interface to MINOS errors from Minuit or NewMinuit (29 June 2009)] * [EBL Attenuation models (20 June 2009)|#EBL Attenuation models (20 June 2009)] * [SummedLikelihood (11 June 2009)|#SummedLikelihood. Unbinned and binned likelihood analysis objects can now be combined in joint analyses. (11 June 2009)] * [SmoothBrokenPowerLaw (11 June 2009)|#SmoothBrokenPowerLaw (11 June 2009)] * [Flux, energy flux, and upper limit calculations for diffuse sources (22 Feb 2009)|#Flux, energy flux, and upper limit calculations for diffuse sources (22 Feb 2009)] * [Diffuse response calculations and FITS image templates (22 Feb 2009)|#Diffuse response calculations and FITS image templates (22 Feb 2009)] * [Analysis object creation using the hoops/ape interface|#Analysis object creation using the hoops/ape interface] * [Photon and Energy Flux Calculations|#Photon and Energy Flux Calculations] h2. Calculating SEDs directly from pyLikelihood (8 November 2011) SEDs can be computed directly from pyLikelihod using a new SED class. This code has the advantage that it does not create any temporary files and does not require multiple gtlike jobs or xml files. The only additional dependency of the code is scipy. Usage example: {code} # SEDs can only be computed for binned analysis. # First, created the BinnedAnalysis object... like=BinnedAnalysis(...) ... # Now created the SED object # This object is in pyLikelihood from SED import SED name='Vela' sed = SED(like,name) # There are many helpful flags to modify how the SED is created sed = SED(like,name, # minimum TS in which to compute an upper limit min_ts=4, # Upper limit algoirthm: bayesian or frequentist ul_algorithm='bayesian', # confidence level of upper limtis ul_confidence=.95, # By default, the energy binning is consistent with # the binning in the ft1 file. This can be modified, but the # new bins must line up with the ft1 file's bin edges bin_edges = [1e2, 1e3, 1e4, 1e4]) # save data points to a file sed.save('sed_Vela.dat') # The SED can be plotted sed.plot('sed_Vela.png') # And finally, the results of the SED fit can be packaged # into a convenient dictionary for further use results=sed.todict() # To load the data points into a new python script: results_dictionary=eval(open('sed_vela.dat').read()) {code} h2. Prior distributions on fit parameters (4 February 2011) |
Prior distributions on fit parameters (4 February 2011)
optimizers-02-21-00,
...
Likelihood-17-05-01,
...
pyLikelihood-01-27-01
...
1D
...
priors
...
can
...
now
...
be
...
assigned
...
to
...
individual
...
fit
...
parameters
...
in
...
the
...
Python
...
interface.
...
The
...
current
...
implementation
...
allows
...
for
...
Gaussian
...
pdfs,
...
but
...
the
...
interface
...
can
...
also
...
accomodate
...
other
...
pdfs:
...
the
...
corresponding
...
Function
...
classes
...
just
...
need
...
to
...
be
...
implemented.
...
Usage
...
examples:
| Code Block |
|---|
} >>> like[i].addGaussianPrior(-2.3, 0.1) # Add a Gaussian prior to parameter i with a mean value of -2.3 and root-variance (sigma) of 0.1 >>> like[i].addPrior('LogGaussian') # Add a prior function by name. Default parameter values will be used. >>> print like[i].getPriorParams() # This returns a dictionary of the parameters of the function representing the pdf. {'Sigma': 0.01, 'Norm': 1.0, 'Mean':-2.29999999999999998} >>> like[i].setGaussianPriorParams(-2.3, 0.01) # Set the prior parameter values (mean, sigma) for the Gaussian case. >>> like[i].setPriorParams(Mean=-2.3) # Set the prior parameter values by name for any prior function. {code} h2. User defined error and contour level in Minos (3 May 2010) |
User defined error and contour level in Minos (3 May 2010)
optimizers-02-18-00
...
In
...
MINUIT
...
parlance,
...
the
...
parameter
...
UP
...
defines
...
the
...
error
...
level
...
returned
...
by
...
the
...
routines
...
MIGRAD/HESSE/MINOS.
...
If
...
FCN
...
is
...
the
...
function
...
to
...
be
...
minimized,
...
the
...
returned
...
error
...
is
...
this
...
value
...
which,
...
added
...
to
...
the
...
best
...
fit
...
value
...
of
...
the
...
parameter,
...
increases
...
FCN
...
by
...
an
...
amount
...
of
...
UP.
...
If
...
FCN
...
is
...
the
...
usual
...
chi-square
...
function
...
(a.k.a
...
weighted
...
least
...
square
...
function),
...
UP=1
...
yields
...
the
...
usual
...
1-sigma
...
error.
...
If
...
FCN
...
is
...
minus
...
the
...
log
...
Likelihood
...
function,
...
then
...
UP=0.5
...
yields
...
the
...
1-sigma
...
error.
...
In
...
the
...
ScienceTools,
...
the
...
optimizers
...
package
...
defines
...
the
...
FCN
...
as
...
minus
...
the
...
log
...
likelihood
...
function.
...
The
...
new
...
implementation
...
allow
...
the
...
user
...
to
...
define
...
the
...
error
...
level
...
he
...
wish
...
to
...
obtain
...
from
...
MINUIT,
...
via
...
the
...
"level"
...
argument,
...
which
...
is
...
defined
...
as
...
:
...
UP=level/2.
...
It
...
is
...
the
...
responsibility
...
of
...
the
...
user
...
to
...
understand
...
what
...
"level"
...
should
...
be,
...
depending
...
on
...
his
...
use
...
case.
...
What
...
the
...
code
...
implementation
...
ensures
...
is
...
the
...
following
...
:
...
"level"
...
is
...
that
...
value
...
of
...
the
...
function
...
-2DeltaLogLikelihood(p)
...
that,
...
when
...
reached
...
for
...
a
...
parameter
...
value
...
p,
...
will
...
drive
...
MINUIT
...
to
...
return
...
as
...
the
...
error
...
the
...
value
...
(p-p_min),
...
where
...
p_min
...
is
...
the
...
best
...
fit
...
value
...
of
...
the
...
parameter
...
p.
...
Under
...
the
...
assumption
...
that
...
-2DeltaLogLikelihood
...
is
...
distributed
...
as
...
the
...
chi-square
...
probability
...
distribution
...
with
...
one
...
degree
...
of
...
freedom,
...
level=4
...
with
...
thus
...
yield
...
two-sided
...
2-sigma
...
errors.
...
This
...
is
...
because
...
there
...
is
...
5%
...
of
...
the
...
total
...
chi2
...
density
...
beyond
...
the
...
value
...
4
...
(actually
...
~3.85),
...
which
...
translates
...
into
...
2-sigma
...
errors,
...
as
...
there
...
is
...
2.5%
...
of
...
the
...
Normal
...
probability
...
density
...
beyond
...
2
...
(actually
...
~2.3%),
...
and
...
we
...
are
...
looking
...
at
...
2-sided
...
confidence
...
intervals.
...
Finally,
...
the
...
MNCONTOUR
...
contour
...
analysis
...
from
...
MINUIT,
...
which
...
essentially
...
runs
...
MINOS
...
on
...
a
...
pair
...
of
...
parameters,
...
is
...
now
...
also
...
exposed.
...
This
...
needs
...
further
...
testing,
...
and
...
API
...
improvement
...
so
...
that
...
the
...
resulting
...
contour
...
points
...
can
...
be
...
returned
...
within
...
python
...
:
...
they
...
are
...
currently
...
just
...
printed
...
out
...
onscreen,
...
together
...
with
...
the
...
contour
...
(ascii
...
plotting
...
like
...
in
...
the
...
old
...
days
...
:)
...
).
...
Take
...
note
...
that
...
these
...
are
...
now
...
2-parameter
...
errors,
...
so
...
that
...
level=1
...
is
...
not
...
a
...
1-sigma
...
contour
...
(though
...
you
...
do
...
get
...
the
...
1-sigma
...
one
...
parameter
...
errors
...
by
...
projecting
...
the
...
extrema
...
of
...
the
...
contour
...
along
...
the
...
corresponding
...
axis).
...
For
...
more
...
information
...
on
...
MINUIT?
...
please
...
look
...
at
...
...
...
...
...
...
...
...
,
...
and
...
the
...
...
...
...
...
Composite2 (11 July 2010)
Likelihood-16-10-00,
...
pyLikelihood-01-25-00
...
This
...
is
...
similar
...
to
...
the
...
CompositeLikelihod
...
and
...
SummedLikelihood
...
classes
...
(all
...
three
...
are
...
accessible
...
via
...
the
...
pyLikelihood
...
interface)
...
in
...
that
...
log-likelihood
...
objects
...
may
...
be
...
combined
...
to
...
form
...
a
...
single
...
objective
...
function.
...
Composite2
...
allows
...
users
...
to
...
tie
...
together
...
arbitrary
...
sets
...
of
...
parameter
...
across
...
(and
...
within)
...
the
...
different
...
log-likelihood
...
components
...
of
...
the
...
composite.
...
It
...
can
...
therefore
...
reproduce
...
the
...
functionality
...
of
...
the
...
CompositeLikelihood
...
model,
...
where
...
the
...
parameters,
...
except
...
for
...
the
...
normalizations,
...
for
...
the
...
"stacked"
...
sources
...
are
...
tied
...
together;
...
it
...
can
...
also
...
reproduce
...
the
...
functionality
...
of
...
the
...
SummedLikelihood,
...
where
...
all
...
of
...
the
...
corresponding
...
parameters
...
for
...
models
...
of
...
the
...
different
...
components
...
are
...
tied
...
together.
...
The
...
SummedLikelihood
...
implementation
...
is
...
still
...
preferred
...
for
...
that
...
application
...
since
...
it
...
supports
...
the
...
UpperLimits.py
...
and
...
IntegralUpperLimit.py
...
modules
...
whereas
...
Composite2
...
does
...
not.
...
Composite2
...
does
...
support
...
the
...
minosError
...
function
...
that
...
is
...
also
...
available
...
from
...
CompositeLikelihood.
...
Composite2
...
has
...
not
...
been
...
extensively
...
tested.
...
Here
...
are
...
scripts
...
that
...
demonstrate
...
the
...
Composite2
...
equivalence
...
to
...
...
and
...
...
.
...
For
...
clarity,
...
here
...
is
...
the
...
...
...
...
used
...
with
...
those
...
scripts.
...
RadialProfile
...
model
...
(5
...
July
...
2010)
...
Likelihood-16-
...
08-02
Similar to the RadialSource model in celestialSource/genericSources,
...
this
...
model
...
defines
...
an
...
azimuthally
...
symmetric
...
diffuse
...
source
...
that
...
is
...
specified
...
by
...
an
...
ascii
...
file
...
containing
...
the
...
radial
...
profile
...
as
...
a
...
function
...
of
...
off-source
...
angle.
| Code Block | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| =
|
|
|
|
| |||||||
} <source name="test_profile" type="DiffuseSource"> <spectrum type="PowerLaw"> <parameter free="1" max="1000.0" min="0.001" name="Prefactor" scale="1e-09" value="10.0"/> <parameter free="1" max="-1.0" min="-5.0" name="Index" scale="1.0" value="-2.0"/> <parameter free="0" max="2000.0" min="30.0" name="Scale" scale="1.0" value="100.0"/> </spectrum> <spatialModel type="RadialProfile" file="radial_profile.txt"> <parameter free="0" max="10" min="0" name="Normalization" scale="1.0" value="1"/> <parameter free="0" max="360." min="-360" name="RA" scale="1.0" value="82.74"/> <parameter free="0" max="90" min="-90" name="DEC" scale="1.0" value="13.38"/> </spatialModel> </source> {code} An [example template file|^radial_profile.txt] for the radial profile is attached. The first column is the offset angle in degrees from the source center (given by RA, DEC in the xml model def), and the second column is the differential intensity. The units of the function in that file times the spectral part of the xml model definition should be |
An example template file for the radial profile is attached. The first column is the offset angle in degrees from the source center (given by RA, DEC in the xml model def), and the second column is the differential intensity. The units of the function in that file times the spectral part of the xml model definition should be photons/cm^2/s/sr,
...
just
...
as
...
for
...
any
...
other
...
diffuse
...
source.
...
NB:
...
Linear
...
interpolation
...
is
...
used
...
for
...
evaluating
...
the
...
profile
...
using
...
the
...
points
...
in
...
the
...
template
...
file,
...
and
...
the
...
code
...
will
...
simply
...
try
...
to
...
extrapolate
...
beyond
...
the
...
first
...
and
...
last
...
entries.
...
So
...
the
...
first
...
point
...
should
...
be
...
at
...
offset
...
zero,
...
and
...
the
...
last
...
two
...
entries
...
should
...
both
...
be
...
set
...
to
...
zero
...
intensity,
...
otherwise
...
the
...
extrapolation
...
can
...
go
...
negative
...
at
...
large
...
angles.
...
This
...
source
...
has
...
not
...
been
...
extensively
...
tested.
...
Upper
...
limits
...
using
...
the
...
(semi-)Bayesian
...
method
...
of
...
Helene
...
(21
...
September
...
2009)
...
pyLikelihood
...
v1r17
...
For
...
background,
...
see
...
...
...
...
to
...
the
...
Catalog
...
group.
...
This
...
functionality
...
is
...
now
...
available
...
in
...
two
...
ways:
...
as
...
a
...
new
...
method,
...
bayesianUL(...),
...
in
...
the
...
existing
...
UpperLimit
...
class,
...
and
...
as
...
a
...
separate
...
function
...
in
...
the
...
IntegralUpperLimit.py
...
module.
...
The
...
latter
...
was
...
implemented
...
by
...
Stephen
...
using
...
scipy
...
(which
...
we
...
do
...
not
...
distribute
...
with
...
ST
...
python).
...
The
...
bayesianUL(...)
...
method
...
does
...
not
...
require
...
scipy
...
and
...
can
...
only
...
be
...
run
...
after
...
UpperLimit.compute(...)
...
is
...
run
...
first.
...
The
...
scipy
...
version
...
is
...
more
...
robust
...
in
...
some
...
cases;
...
the
...
bayesianUL(...)
...
function
...
can
...
return
...
a
...
spuriously
...
low
...
upper
...
limit
...
if
...
the
...
pyLikelihood
...
object
...
is
...
not
...
initially
...
at
...
a
...
good
...
minimum.
...
Example
...
of
...
usage:
| Code Block | ||||
|---|---|---|---|---|
| =
| |||
} from UnbinnedAnalysis import * import UpperLimits import IntegralUpperLimit # This requires scipy like = unbinnedAnalysis(evfile='filtered.fits', scfile='test_scData_0000.fits', expmap='expMap.fits', expcube='expCube.fits', srcmdl='srcModel.xml', irfs='P6_V3_DIFFUSE', optimizer='MINUIT') src = 'my_source' emin = 100 emax = 3e5 PhIndex = like.par_index(src, 'Index') like[PhIndex] = -2.2 like.freeze(PhIndex) like.fit(0) ul = UpperLimits.UpperLimits(like) ul_prof, par_prof = ul[src].compute(emin=emin, emax=emax, delta=2.71/2) print "Upper limit using profile method: ", ul_prof ul_bayes, par_bayes = ul[src].bayesianUL(emin=emin, emax=emax, cl=0.95) print "Using default implementation of method of Helene: ", ul_bayes ul_scipy, results_scipy = IntegralUpperLimit.calc(like, src, ul=0.95, emin=emin, emax=emax) print "Using S. Fegan's module (requiring scipy): ", ul_scipy {code} |
Output
...
(note
...
use
...
of
...
the
...
local
...
ipython,
...
which
...
has
...
scipy
...
installed):
| No Format |
|---|
} ki-rh2[jchiang] which ipython /afs/slac/g/ki/software/python/2.5.1/i386_linux26/bin/ipython ki-rh2[jchiang] ipython analysis.py 0 0.005535583927 2.21673610667e-06 8.05446445813e-07 1 0.00700983628826 0.0713089268528 1.01995522036e-06 2 0.00848408864951 0.24764020254 1.23446399491e-06 3 0.00995834101077 0.49775117816 1.44897276946e-06 4 0.011432593372 0.802803851635 1.66348154402e-06 5 0.0135311686428 1.30808036117 1.96883144303e-06 6 0.0139522586249 1.417706337 2.03010147956e-06 Upper limit using profile method: 1.99505486283e-06 Using default implementation of method of Helene: 2.29836280879e-06 Using S. Fegan's module (requiring scipy): 2.31242668344e-06 Python 2.5.1 (r251:54863, May 23 2007, 23:16:17) Type "copyright", "credits" or "license" for more information. IPython 0.8.1 -- An enhanced Interactive Python. ? -> Introduction to IPython's features. %magic -> Information about IPython's 'magic' % functions. help -> Python's own help system. object? -> Details about 'object'. ?object also works, ?? prints more. In [1]: {noformat} |
As
...
usual,
...
the
...
units
...
of
...
the
...
flux
...
upper
...
limits
...
are
...
photons/cm^2/s.
...
Interface
...
to
...
MINOS
...
errors
...
from
...
Minuit
...
or
...
NewMinuit
...
(29
...
June
...
2009)
...
optimizers
...
v2r17p1,
...
pyLikelihood
...
v1r15
...
The
...
Minuit
...
and
...
Minuit2
...
(NewMinuit)
...
libraries
...
can
...
calculate
...
asymmetric
...
errors
...
that
...
are
...
based
...
on
...
the
...
profile
...
likelihood
...
method.
...
I've
...
modified
...
the
...
python
...
interface
...
in
...
pyLikelihood
...
to
...
make
...
it
...
easier
...
to
...
have
...
the
...
code
...
evaluate
...
these
...
errors.
...
Note
...
that
...
these
...
errors
...
are
...
only
...
available
...
for
...
the
...
'Minuit'
...
and
...
'NewMinuit'
...
optimizers.
...
Here
...
is
...
a
...
script
...
illustrating
...
the
...
interface.
...
Parameters
...
can
...
be
...
specified
...
by
...
index
...
number
...
(as
...
displayed
...
by
...
...
like.model")
...
or
...
by
...
source
...
name
...
and
...
parameter
...
name:
| Code Block |
|---|
} from UnbinnedAnalysis import * like = unbinnedAnalysis(mode='h', optimizer='NewMinuit', evfile='ft1_03.fits', scfile='FT2.fits', srcmdl='3c273_03.xml', expmap='expMap_03_P6_V3.fits', expcube='expCube_03_P6_V3.fits', irfs='P6_V3_DIFFUSE') try: print like.minosError('3C 273', 'Integral') except RuntimeError, message: print "Caught a RuntimeError: " print message like.fit(0) print like.model print "Minos errors:" print "3C 273, Integral: ", like.minosError('3C 273', 'Integral') print "3C 273, Integral: ", like.minosError(0) print "3C 273, Index: ", like.minosError('3C 273', 'Index') try: print "ASO0267, Integral:", like.minosError('ASO0267', 'Integral') except RuntimeError: par_index = like.par_index('ASO0267', 'Integral') print "ASO0267, Integral parameter index: ", par_index print "Parabolic error estimate: ", like[par_index].error() {code} |
The
...
function
...
'like.par_index(...)'
...
is
...
also
...
new.
...
It
...
returns
...
the
...
index
...
of
...
the
...
parameter
...
when
...
given
...
the
...
source
...
and
...
parameter
...
names.
...
Here
...
is
...
the
...
output:
| No Format |
|---|
} Caught a RuntimeError: To evaluate minos errors, a fit must first be performed using the Minuit or NewMinuit optimizers. <...skip some unsupressable Minuit2 output...> 3C 273 Spectrum: PowerLaw2 0 Integral: 7.343e-01 2.525e-01 1.000e-05 1.000e+03 ( 1.000e-06) 1 Index: -2.592e+00 2.824e-01 -5.000e+00 -1.000e+00 ( 1.000e+00) 2 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 2.000e+05 ( 1.000e+00) fixed 3 UpperLimit: 3.000e+05 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 3C 279 Spectrum: PowerLaw2 4 Integral: 1.241e+00 4.307e-01 1.000e-05 1.000e+03 ( 1.000e-06) 5 Index: -2.981e+00 3.197e-01 -5.000e+00 -1.000e+00 ( 1.000e+00) 6 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 2.000e+05 ( 1.000e+00) fixed 7 UpperLimit: 3.000e+05 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed ASO0267 Spectrum: PowerLaw2 8 Integral: 1.886e-01 4.058e-01 1.000e-05 1.000e+03 ( 1.000e-06) 9 Index: -5.000e+00 7.679e-02 -5.000e+00 -1.000e+00 ( 1.000e+00) 10 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 2.000e+05 ( 1.000e+00) fixed 11 UpperLimit: 3.000e+05 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed Extragalactic Diffuse Spectrum: PowerLaw 12 Prefactor: 3.750e+00 1.052e+00 1.000e-05 1.000e+02 ( 1.000e-07) 13 Index: -2.358e+00 7.992e-02 -3.500e+00 -1.000e+00 ( 1.000e+00) 14 Scale: 1.000e+02 0.000e+00 5.000e+01 2.000e+02 ( 1.000e+00) fixed GalProp Diffuse Spectrum: ConstantValue 15 Value: 3.474e-01 7.606e-01 0.000e+00 1.000e+01 ( 1.000e+00) Minos errors: 3C 273, Integral: (-0.21305533965274412, 0.30406941767501522) 3C 273, Integral: (-0.21305533965275783, 0.30406941767466594) 3C 273, Index: (-0.30561922642904105, 0.26192991818914718) Attempt to set the value outside of existing bounds. Value -0.217096 is not between1e-05 and 1000 ASO0267, Integral: Minos error encountered for parameter 8. Attempting to reset free parameters. ASO0267, Integral parameter index: 8 Parabolic error estimate: 0.405782121649 ki-rh2[jchiang] {noformat} |
The
...
Minos
...
implementation
...
in
...
'NewMinuit'
...
tends
...
to
...
work
...
better
...
than
...
that
...
in
...
'Minuit'.
...
PS
...
:
...
As
...
can
...
be
...
seen
...
in
...
the
...
example
...
above,
...
if
...
Minuit2
...
(NewMinuit)
...
needs
...
to
...
scan
...
the
...
parameter
...
value
...
outside
...
the
...
user-specified
...
bounds,
...
the
...
code
...
will
...
generate
...
an
...
exception.
...
However,
...
Minuit2
...
can
...
still
...
fail
...
to
...
find
...
an
...
upper
...
or
...
lower
...
value
...
that
...
corresponds
...
to
...
the
...
desired
...
change
...
in
...
log-likelihood
...
(+0.5
...
for
...
1-sigma
...
errors)
...
even
...
if
...
it
...
doesn't
...
try
...
to
...
set
...
the
...
parameter
...
outside
...
of
...
the
...
valid
...
range.
...
In
...
this
...
case,
...
Minuit2
...
appears
...
to
...
give
...
up
...
and
...
just
...
return
...
the
...
parabolic
...
error
...
estimate:
| No Format |
|---|
} >>> print like['my_source'] my_source Spectrum: PowerLaw2 28 Integral: 4.224e-02 2.188e-03 0.000e+00 1.000e+03 ( 1.000e-06) 29 Index: -1.628e+00 2.291e-02 -5.000e+00 0.000e+00 ( 1.000e+00) 30 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 31 UpperLimit: 1.000e+05 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed >>> like.minosError('my_source', 'Integral') Info in <Minuit2>: VariableMetricBuilder: no improvement in line search Info in <Minuit2>: VariableMetricBuilder: iterations finish without convergence. Info in <Minuit2>: VariableMetricBuilder : edm = 0.0465751 Info in <Minuit2>: requested : edmval = 1.25e-05 Info in <Minuit2>: MnMinos could not find Upper Value for Parameter : par = 15 Info in <Minuit2>: VariableMetricBuilder: matrix not pos.def, gdel > 0 Info in <Minuit2>: gdel = 0.000345418 Info in <Minuit2>: negative or zero diagonal element in covariance matrix : i = 8 Info in <Minuit2>: added to diagonal of Error matrix a value : dg = 792.328 Info in <Minuit2>: gdel = -88.8478 Info in <Minuit2>: VariableMetricBuilder: no improvement in line search Info in <Minuit2>: VariableMetricBuilder: iterations finish without convergence. Info in <Minuit2>: VariableMetricBuilder : edm = 216.425 Info in <Minuit2>: requested : edmval = 1.25e-05 Info in <Minuit2>: MnMinos could not find Lower Value for Parameter : par = 15 Out[26]: (-0.0021882980216484244, 0.0021882980216484244) >>> {noformat} |
The
...
"Info"
...
text
...
from
...
Minuit2
...
is
...
not
...
trappable
...
by
...
the
...
client
...
code,
...
so
...
currently
...
there
...
is
...
no
...
way
...
for
...
pyLikelihood
...
to
...
determine
...
definitively
...
that
...
Minuit2::MnMinos
...
has
...
given
...
up.
...
One
...
workaround
...
would
...
be
...
to
...
test
...
if
...
the
...
minos
...
error
...
is
...
the
...
same
...
as
...
the
...
parabolic
...
estimate.
...
EBL
...
Attenuation
...
models
...
(20
...
June
...
2009)
...
celestialSources/eblAtten
...
v0r7p1,
...
optimizers
...
v2r17,
...
Likelihood
...
v15r2
...
I've
...
added
...
access
...
to
...
the
...
EBL
...
attenuation
...
models
...
that
...
are
...
available
...
to
...
gtobssim
...
(and
...
Gleam)
...
simulations
...
via
...
the
...
...
package.
...
Here
...
is
...
an
...
example
...
xml
...
definition
...
for
...
the
...
PowerLaw2
...
spectral
...
model:
| Code Block |
|---|
} <spectrum type="EblAtten::PowerLaw2"> <parameter free="1" max="1000.0" min="1e-05" name="Integral" scale="1e-06" value="2.0"/> <parameter free="1" max="-1.0" min="-5.0" name="Index" scale="1.0" value="-2.0"/> <parameter free="0" max="200000.0" min="20.0" name="LowerLimit" scale="1.0" value="20.0"/> <parameter free="0" max="200000.0" min="20.0" name="UpperLimit" scale="1.0" value="2e5"/> <parameter free="1" max="10" min="0" name="tau_norm" scale="1.0" value="1"/> <parameter free="0" max="10" min="0" name="redshift" scale="1.0" value="0.5"/> <parameter free="0" max="8" min="0" name="ebl_model" scale="1.0" value="0"/> </spectrum> {code} |
As
...
seen
...
in
...
this
...
example,
...
three
...
parameters
...
are
...
added
...
to
...
the
...
underlying
...
spectral
...
model:
...
tau_norm,
...
redshift,
...
and
...
ebl_model.
...
The
...
models
...
with
...
EBL
...
attenuation
...
that
...
are
...
available
...
are
...
EblAtten::PowerLaw2 |
...
EblAtten::BrokenPowerLaw2 |
...
EblAtten::LogParabola |
...
EblAtten::BandFunction |
...
EblAtten::SmoothBrokenPowerLaw |
...
EblAtten::FileFunction |
...
EblAtten::ExpCutoff |
...
EblAtten::BPLExpCutoff |
...
EblAtten::PLSuperExpCutoff |
...
The
...
current
...
list
...
of
...
EBL
...
models
...
are
...
model |
|---|
...
id |
|---|
...
model name | |
|---|---|
0 | Kneiske |
1 | Primack05 |
2 | Kneiske_HighUV |
3 | Stecker05 |
4 | Franceschini |
5 | Finke |
6 | Gilmore |
The definitive list can be found in the enums defined in the EblAtten class, and references for each model are given in the header comments for the implementation of the optical depth functions.
SummedLikelihood. Unbinned and binned likelihood analysis objects can now be combined in joint analyses. (11 June 2009)
Likelihood v15r0p3, pyLikelihood v1r14p1
Here is a script that does a joint analysis of unbinned likelihood with binned front and binned back event likelihoods:
| Code Block |
|---|
name || | 0 | Kneiske | | 1 | Primack05 | | 2 | Kneiske_HighUV | | 3 | Stecker05 | | 4 | Franceschini | | 5 | Finke | | 6 | Gilmore | The definitive list can be found in the enums defined in the [EblAtten|http://www-glast.stanford.edu/cgi-bin/viewcvs/celestialSources/eblAtten/eblAtten/EblAtten.h?view=markup] class, and references for each model are given in the header comments for the [implementation of the optical depth functions|http://www-glast.stanford.edu/cgi-bin/viewcvs/celestialSources/eblAtten/src/IRB_routines.cxx?view=markup]. h2. SummedLikelihood. Unbinned and binned likelihood analysis objects can now be combined in joint analyses. (11 June 2009) Likelihood v15r0p3, pyLikelihood v1r14p1 Here is a script that does a joint analysis of unbinned likelihood with binned front and binned back event likelihoods: {code} from UnbinnedAnalysis import * from BinnedAnalysis import * from SummedLikelihood import * from UpperLimits import * scfile = 'test_scData_0000.fits' ltcube = 'expCube.fits' irfs = 'P6_V3_DIFFUSE' optimizer = 'MINUIT' srcmdl = 'anticenter_model.xml' like = unbinnedAnalysis(evfile='filtered.fits', scfile=scfile, expmap='expMap.fits', expcube=ltcube, irfs=irfs, optimizer=optimizer, srcmdl=srcmdl) like_u = unbinnedAnalysis(evfile='filtered_gt_3GeV.fits', scfile=scfile, expmap='expMap.fits', expcube=ltcube, irfs=irfs, optimizer=optimizer, srcmdl=srcmdl) like_f = binnedAnalysis(irfs=irfs, expcube=ltcube, srcmdl=srcmdl, optimizer=optimizer, cmap='smaps_front_3GeV.fits', bexpmap='binned_exp_front_lt_3GeV.fits') like_b = binnedAnalysis(irfs=irfs, expcube=ltcube, srcmdl=srcmdl, optimizer=optimizer, cmap='smaps_back_3GeV.fits', bexpmap='binned_exp_back_lt_3GeV.fits') summed_like = SummedLikelihood() summed_like.addComponent(like_u) summed_like.addComponent(like_f) summed_like.addComponent(like_b) like.fit(0) print "Unbinned results:" print like.model summed_like.fit(0) print "Combined unbinned and binned results:" print summed_like.model ul = UpperLimits(like) ul_summed = UpperLimits(summed_like) print "Unbinned upper limit example:" print ul['PKS 0528+134'].compute() print "Combined unbinned and binned upper limit example:" print ul_summed['PKS 0528+134'].compute() print like.Ts('Crab') print summed_like.Ts('Crab') {code} |
Here
...
is
...
the
...
output:
| No Format |
|---|
} Unbinned results: Crab Spectrum: PowerLaw2 0 Integral: 1.955e+01 3.198e+00 1.000e-05 1.000e+03 ( 1.000e-07) 1 Index: -2.039e+00 1.091e-01 -5.000e+00 0.000e+00 ( 1.000e+00) 2 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 3 UpperLimit: 5.000e+05 0.000e+00 2.000e+01 5.000e+05 ( 1.000e+00) fixed Extragalactic Diffuse Spectrum: PowerLaw 4 Prefactor: 1.049e+00 5.228e-01 1.000e-05 1.000e+02 ( 1.000e-07) 5 Index: -2.136e+00 1.772e-01 -3.500e+00 -1.000e+00 ( 1.000e+00) 6 Scale: 1.000e+02 0.000e+00 5.000e+01 2.000e+02 ( 1.000e+00) fixed GalProp Diffuse Spectrum: ConstantValue 7 Value: 1.229e+00 6.405e-02 0.000e+00 1.000e+01 ( 1.000e+00) Geminga Spectrum: PowerLaw2 8 Integral: 3.463e+01 3.295e+00 1.000e-05 1.000e+03 ( 1.000e-07) 9 Index: -1.700e+00 5.405e-02 -5.000e+00 0.000e+00 ( 1.000e+00) 10 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 11 UpperLimit: 5.000e+05 0.000e+00 2.000e+01 5.000e+05 ( 1.000e+00) fixed PKS 0528+134 Spectrum: PowerLaw2 12 Integral: 1.214e+01 3.081e+00 1.000e-05 1.000e+03 ( 1.000e-07) 13 Index: -2.555e+00 2.340e-01 -5.000e+00 0.000e+00 ( 1.000e+00) 14 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 15 UpperLimit: 5.000e+05 0.000e+00 2.000e+01 5.000e+05 ( 1.000e+00) fixed Combined unbinned and binned results: Crab Spectrum: PowerLaw2 0 Integral: 1.927e+01 3.175e+00 1.000e-05 1.000e+03 ( 1.000e-07) 1 Index: -2.026e+00 1.083e-01 -5.000e+00 0.000e+00 ( 1.000e+00) 2 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 3 UpperLimit: 5.000e+05 0.000e+00 2.000e+01 5.000e+05 ( 1.000e+00) fixed Extragalactic Diffuse Spectrum: PowerLaw 4 Prefactor: 7.895e-01 6.264e-01 1.000e-05 1.000e+02 ( 1.000e-07) 5 Index: -2.088e+00 2.110e-01 -3.500e+00 -1.000e+00 ( 1.000e+00) 6 Scale: 1.000e+02 0.000e+00 5.000e+01 2.000e+02 ( 1.000e+00) fixed GalProp Diffuse Spectrum: ConstantValue 7 Value: 1.269e+00 7.695e-02 0.000e+00 1.000e+01 ( 1.000e+00) Geminga Spectrum: PowerLaw2 8 Integral: 3.494e+01 3.327e+00 1.000e-05 1.000e+03 ( 1.000e-07) 9 Index: -1.701e+00 5.386e-02 -5.000e+00 0.000e+00 ( 1.000e+00) 10 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 11 UpperLimit: 5.000e+05 0.000e+00 2.000e+01 5.000e+05 ( 1.000e+00) fixed PKS 0528+134 Spectrum: PowerLaw2 12 Integral: 1.176e+01 3.008e+00 1.000e-05 1.000e+03 ( 1.000e-07) 13 Index: -2.534e+00 2.311e-01 -5.000e+00 0.000e+00 ( 1.000e+00) 14 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 15 UpperLimit: 5.000e+05 0.000e+00 2.000e+01 5.000e+05 ( 1.000e+00) fixed Unbinned upper limit example: 0 12.1361746852 -0.000307310165226 1.21960706796e-06 1 13.368492501 0.0833717813548 1.34368135193e-06 2 14.6008103168 0.305775613851 1.46778865044e-06 3 15.8331281326 0.646238077657 1.59193260068e-06 4 17.0654459483 1.08809170817 1.71611295547e-06 5 17.8801837016 1.42911471373 1.7982332395e-06 (1.7803859938766175e-06, 17.703116306269838) Combined unbinned and binned upper limit example: 0 11.7635006963 -0.000623093163085 1.18207829753e-06 1 12.9668988172 0.0780380520519 1.30323312058e-06 2 14.1702969382 0.291922864584 1.42440474102e-06 3 15.3736950591 0.621846241655 1.54561984399e-06 4 16.57709318 1.05134415647 1.66686579099e-06 5 17.7804913009 1.56725279011 1.78814776158e-06 (1.7382504833201744e-06, 17.285394704120577) 282.100179467 271.963534683 {noformat} |
It
...
would
...
be
...
wise
...
to
...
ensure
...
that
...
the
...
datasets
...
used
...
in
...
the
...
summed
...
likelihoods
...
do
...
not
...
intersect.
...
SmoothBrokenPowerLaw
...
(11
...
June
...
2009)
...
Likelihood
...
v15r1p2
...
Benoit
...
implemented
...
and
...
added
...
a
...
new
...
spectral
...
model.
...
Documentation
...
can
...
be
...
found
...
in
...
the
...
workbook
...
here.
Flux, energy flux, and upper limit calculations for diffuse sources (22 Feb 2009)
pyLikelihood v1r10p1; Likelihood v14r4
Flux, energy flux and upper limit calculations can now be made for diffuse sources in the python interface:
| Code Block |
|---|
|http://glast-ground.slac.stanford.edu/workbook/pages/sciTools_likelihoodTutorial/likelihoodTutorial.htm#createSourceModel]. h2. Flux, energy flux, and upper limit calculations for diffuse sources (22 Feb 2009) pyLikelihood v1r10p1; Likelihood v14r4 Flux, energy flux and upper limit calculations can now be made for diffuse sources in the python interface: {code} >>> like.model Extragalactic Diffuse Spectrum: PowerLaw 0 Prefactor: 1.450e+00 4.286e-01 1.000e-05 1.000e+02 ( 1.000e-07) 1 Index: -2.054e+00 1.366e-01 -3.500e+00 -1.000e+00 ( 1.000e+00) 2 Scale: 1.000e+02 0.000e+00 5.000e+01 2.000e+02 ( 1.000e+00) fixed SNR_map Spectrum: PowerLaw2 3 Integral: 6.953e+03 2.566e+03 0.000e+00 1.000e+10 ( 1.000e-06) 4 Index: -2.039e+00 1.332e-01 -5.000e+00 -1.000e+00 ( 1.000e+00) 5 LowerLimit: 2.000e+01 0.000e+00 2.000e+01 2.000e+05 ( 1.000e+00) fixed 6 UpperLimit: 5.000e+05 0.000e+00 2.000e+01 5.000e+05 ( 1.000e+00) fixed >>> like.flux('Extragalactic Diffuse', 100, 5e5) 0.00017279621316112551 >>> like.fluxError('Extragalactic Diffuse', 100, 5e5) 3.32420171199e-05 >>> like.flux('SNR_map', 100, 5e5) 1.3482399819603797e-06 >>> like.fluxError('SNR_map', 100, 5e5) 2.54429802134e-07 >>> from UpperLimits import UpperLimits >>> ul = UpperLimits(like) >>> ul['Extragalactic Diffuse'].compute() 0 1.44990964752 0.000405768508472 0.000172805047298 1 1.62134142735 0.073917905358 0.000185125831974 2 1.79277320719 0.262761447131 0.000197078187151 3 1.96420498702 0.53963174062 0.00020864241284 4 2.13563676685 0.882155849953 0.000219869719057 5 2.33677514646 1.33948412506 0.000232753706737 6 2.37253076663 1.42681888532 0.000234966127989 (0.00023314676502589186, 2.34312748234) >>> ul['SNR_map'].compute() 0 6953.47703963 7.03513949247e-05 1.34891151045e-06 1 7979.85337599 0.072031366891 1.43605234286e-06 2 9006.22971235 0.246899738264 1.51625454475e-06 3 10032.6060487 0.487519676687 1.59037705975e-06 4 11058.9823851 0.770755618154 1.65973696844e-06 5 12629.8902716 1.25421175769 1.75816622567e-06 6 13039.134979 1.38671547284 1.77783091654e-06 (1.7731240676197149e-06, 12941.1800697) {code} |
The
...
compute()
...
command
...
will
...
use
...
profile
...
likelihood
...
to
...
compute
...
the
...
upper
...
limit
...
and
...
will
...
thus
...
scan
...
in
...
normalization
...
parameter
...
value.
...
The
...
screen
...
output
...
comprises
...
the
...
scan
...
values
...
with
...
columns
...
index,
...
parameter
...
value,
...
delta(log-likelihood),
...
flux.
...
The
...
values
...
that
...
are
...
returned
...
are
...
the
...
total
...
flux
...
upper
...
limit
...
(i.e.,
...
integrated
...
over
...
all
...
angles)
...
and
...
the
...
corresponding
...
normalization
...
parameter
...
value.
...
Diffuse
...
response
...
calculations
...
and
...
FITS
...
image
...
templates
...
(22
...
Feb
...
2009)
...
Prior
...
to
...
Likelihood
...
v14r3,
...
there
...
had
...
been
...
a
...
long-standing
...
problem
...
with
...
computing
...
diffuse
...
responses
...
(with
...
gtdiffrsp
...
)
...
for
...
diffuse
...
sources
...
that
...
have
...
FITS
...
image
...
templates
...
for
...
discrete
...
sources,
...
such
...
as
...
SNRs
...
or
...
molecular
...
clouds.
...
In
...
summary,
...
gtdiffrsp
...
performs
...
the
...
following
...
integral
...
for
...
each
...
diffuse
...
source
...
component
...
| Latex |
|---|
i |
| Latex |
|---|
}i{latex} in the xml model definition: {latex} \newcommand{\e}{{\varepsilon}} \newcommand{\ep}{{\varepsilon^\prime}} \newcommand{\phat}{{\hat{p}}} \newcommand{\phatp}{{\hat{p}^\prime}} \begin{equation} \int d\phat S_i(\phat, \ep_j) A(\ep_j, \phat) P(\phatp_j; \ep_j, \phat) \end{equation} {latex} where {latex} |
where
| Latex |
|---|
$\hat{p}_j$ |
...
| Latex |
|---|
$\varepsilon^\prime_j$ |
...
...
the
...
measured
...
direction
...
and
...
measured
...
energy
...
of
...
detected
...
photon
...
| Latex |
|---|
...
$j$ |
...
...
and
...
the
...
integral
...
is
...
performed
...
over
...
(true)
...
directions
...
on
...
the
...
sky
...
| Latex |
|---|
...
$\hat{p}$ |
...
| Latex |
|---|
$A(...)$ |
...
| Latex |
|---|
$P(...)$ |
...
...
the
...
effective
...
area
...
and
...
point
...
spread
...
function,
...
and
...
I
...
have
...
made
...
the
...
approximation
...
equating
...
the
...
true
...
photon
...
energy
...
with
...
the
...
measured
...
value,
...
| Latex |
|---|
...
$\varepsilon = \varepsilon^\prime$ |
...
The problem arises for a discrete diffuse source when its spatial distribution
| Latex |
|---|
$S_i(\hat{p})$ |
...
...
only
...
significantly
...
different
...
from
...
zero
...
far
...
from
...
the
...
location
...
of
...
event
...
| Latex |
|---|
...
$j$ |
...
...
Operationally,
...
the
...
integral
...
is
...
evaluated
...
using
...
an
...
adaptive
...
Romberg
...
integrator
...
that
...
samples
...
the
...
integrand
...
at
...
theta
...
and
...
phi
...
values
...
that
...
are
...
referenced
...
to
...
| Latex |
|---|
...
$\hat{p}_j$ |
...
...
For
...
very
...
compact
...
sources,
...
the
...
integrator
...
will
...
often
...
miss
...
the
...
source
...
entirely
...
and
...
evaluate
...
the
...
integral
...
to
...
zero;
...
and
...
unless
...
the
...
measured
...
photon
...
direction
...
lies
...
directly
...
on
...
a
...
bright
...
part
...
of
...
the
...
extended
...
source,
...
the
...
integral
...
will
...
usually
...
not
...
be
...
very
...
accurate,
...
even
...
if
...
non-zero.
...
In
...
Likelihood
...
v14r3,
...
the
...
code
...
has
...
been
...
modified
...
to
...
restrict
...
the
...
range
...
of
...
theta
...
and
...
phi
...
values
...
over
...
which
...
the
...
integration
...
is
...
performed
...
based
...
on
...
the
...
boundaries
...
of
...
the
...
input
...
map.
...
As
...
long
...
as
...
the
...
input
...
map
...
is
...
not
...
mostly
...
filled
...
with
...
zero
...
or
...
near
...
zero-valued
...
entries,
...
restricting
...
the
...
angular
...
range
...
this
...
way
...
seems
...
to
...
produce
...
accurate
...
diffuse
...
response
...
values.
...
This
...
means
...
that
...
care
...
should
...
be
...
taken
...
in
...
preparing
...
the
...
FITS
...
image
...
templates
...
used
...
for
...
defining
...
diffuse
...
sources
...
in
...
Likelihood.
...
The
...
map
...
should
...
be
...
as
...
small
...
as
...
possible
...
and
...
should
...
exclude
...
regions
...
where
...
the
...
emission
...
is
...
negligible.
...
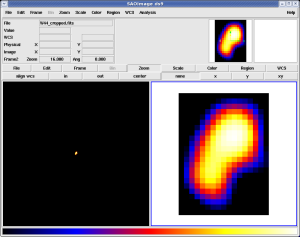
Here
...
are
...
examples
...
of
...
bad
...
(left)
...
and
...
good
...
(right)
...
template
...
files
...
for
...
the
...
same
...
source:
...
The FITS file on the left is 1000x1000 pixels most of which are zero. This is almost optimally bad. Except for events lying right on the source, the diffuse response integrals are almost all zero. The image on the right is the same data, but cropped to the 17x23 pixel region that actually contains relevant data. Performing an unbinned likelihood fit using this smaller map produces accurate results even in the presence of significant Galactic diffuse emission.
Analysis object creation using the hoops/ape interface
The UnbinnedAnalysis and BinnedAnalysis modules contain functions that use the hoops/pil/ape interface to take advantage of the gtlike.par file for specifying inputs. Usage of this interface may be more convenient than creating the UnbinnedObs, UnbinnedAnalysis, BinnedObs, and BinnedAnalysis objects directly. Here are some examples of their use:
- In this example, the call to the unbinnedAnalysis function is made without providing any arguments. The gtlike.par file is read from the user's PFILES path, and the various parameters are prompted for just as when running gtlike (except that the order is a bit different). If one is using the ipython interface with readline enabled, then tab-completion works as well.
Code Block >>> from UnbinnedAnalysis import * >>> like = unbinnedAnalysis() Response functions to use[P6_V1_DIFFUSE] Spacecraft file[test_scData_0000.fits] test_scData_0000.fits Event file[filtered.fits] filtered.fits Unbinned exposure map[none] Exposure hypercube file[expCube.fits] Source model file[anticenter_model.xml] Optimizer (DRMNFB|NEWMINUIT|MINUIT|DRMNGB|LBFGS) [MINUIT] >>> like.model Crab Spectrum: PowerLaw2 0 Integral: 1.540e+01 0.000e+00 1.000e-05 1.000e+03 ( 1.000e-06) 1 Index: -2.190e+00 0.000e+00 -5.000e+00 0.000e+00 ( 1.000e+00) 2 LowerLimit: 2.000e+01 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 3 UpperLimit: 2.000e+05 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed Geminga Spectrum: PowerLaw2 4 Integral: 1.020e+01 0.000e+00 1.000e-05 1.000e+03 ( 1.000e-06) 5 Index: -1.660e+00 0.000e+00 -5.000e+00 0.000e+00 ( 1.000e+00) 6 LowerLimit: 2.000e+01 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 7 UpperLimit: 2.000e+05 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed PKS 0528+134 Spectrum: PowerLaw2 8 Integral: 9.802e+00 0.000e+00 1.000e-05 1.000e+03 ( 1.000e-06) 9 Index: -2.460e+00 0.000e+00 -5.000e+00 0.000e+00 ( 1.000e+00) 10 LowerLimit: 2.000e+01 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed 11 UpperLimit: 2.000e+05 0.000e+00 2.000e+01 3.000e+05 ( 1.000e+00) fixed
...
- Alternatively,
...
- one
...
- can
...
- give
...
- all
...
- of
...
- the
...
- parameters
...
- explicitly.
...
- This
...
- is
...
- useful
...
- for
...
- running
...
- in
...
- scripts.
...
Code Block
...
>>> like2 = unbinnedAnalysis(evfile='filtered.fits', scfile='test_scData_0000.fits', irfs='P6_V1_DIFFUSE', expcube='expCube.fits', srcmdl='anticenter_model.xml', optimizer='minuit', expmap='none')
...
- The mode='h'
...
- option
...
- is
...
- available
...
- as
...
- well.
...
- In
...
- this
...
- case,
...
- one
...
- can
...
- set
...
- a
...
- specific
...
- parameter,
...
- leaving
...
- the
...
- remaining
...
- ones
...
- to
...
- be
...
- read
...
- silently
...
- from
...
- the
...
- gtlike.par
...
- file.
...
Code Block
...
>>> like3 = unbinnedAnalysis(evfile='filtered.fits', mode='h')
...
- A BinnedAnalysis object may be created in a similar fashion using the binnedAnalysis function:
Code Block >>> from BinnedAnalysis import * >>> like = binnedAnalysis() Response functions to use[P6_V1_DIFFUSE::FRONT] Counts map file[smaps_0_20_inc_front.fits] Binned exposure map[binned_expmap_0_20_inc_front.fits] Exposure hypercube file[expCube_0_20_inc.fits] Source model file[Vela_model_0_20_inc_front.xml] Optimizer (DRMNFB|NEWMINUIT|MINUIT|DRMNGB|LBFGS) [MINUIT]
...
Photon and Energy Flux Calculations
With Likelihood v13r18 and pyLikelihood v1r6 (ST v9r8), a facility has been added for calculating photon (ph/cm^2/s)
...
and
...
energy
...
(MeV/cm^2/s)
...
fluxes
...
over
...
a
...
selectable
...
energy
...
range.
...
The
...
errors
...
on
...
these
...
quantities
...
are
...
computed
...
using
...
the
...
procedure
...
described
...
in
...
...
...
made
...
at
...
the
...
...
...
...
...
.
...
Here
...
are
...
some
...
usage
...
examples:
| Code Block |
|---|
} >>> like.model Extragalactic Diffuse Spectrum: PowerLaw 0 Prefactor: 7.527e-02 7.807e-01 1.000e-05 1.000e+02 ( 1.000e-07) 1 Index: -2.421e+00 1.968e+00 -3.500e+00 -1.000e+00 ( 1.000e+00) 2 Scale: 1.000e+02 0.000e+00 5.000e+01 2.000e+02 ( 1.000e+00) fixed GalProp Diffuse Spectrum: ConstantValue 3 Value: 1.186e+00 5.273e-02 0.000e+00 1.000e+01 ( 1.000e+00) Vela Spectrum: BrokenPowerLaw2 4 Integral: 9.176e-02 3.730e-03 1.000e-03 1.000e+03 ( 1.000e-04) 5 Index1: -1.683e+00 5.131e-02 -5.000e+00 -1.000e+00 ( 1.000e+00) 6 Index2: -3.077e+00 2.266e-01 -5.000e+00 -1.000e+00 ( 1.000e+00) 7 BreakValue: 1.716e+03 2.250e+02 3.000e+01 1.000e+04 ( 1.000e+00) 8 LowerLimit: 1.000e+02 0.000e+00 2.000e+01 2.000e+05 ( 1.000e+00) fixed 9 UpperLimit: 3.000e+05 0.000e+00 2.000e+01 5.000e+05 ( 1.000e+00) fixed >>> like.flux('Vela', emin=100, emax=3e5) 9.2005203669330761e-06 >>> like.flux('Vela') 9.2005203669330761e-06 >>> like.fluxError('Vela') 3.73060211564e-07 >>> like.energyFlux('Vela') 0.0047870395747710718 >>> like.energyFluxError('Vela') 0.000260743911378 {code} |
The
...
default
...
energy
...
range
...
for
...
the
...
flux
...
and
...
energy
...
flux
...
calculations
...
is
...
(emin,
...
emax)
...
=
...
(100,
...
3e5)
...
MeV.
...
Either
...
or
...
both
...
of
...
these
...
may
...
be
...
set
...
as
...
keyword
...
arguments
...
to
...
the
...
function
...
call.
...
The
...
errors
...
are
...
available
...
as
...
separate
...
function
...
calls
...
and
...
require
...
that
...
the
...
covariance
...
matrix
...
has
...
been
...
computed
...
using
...
"covar=True"
...
keyword
...
option
...
to
...
the
...
fit
...
function:
| Code Block |
|---|
} >>> like.fit(covar=True) {code} |