Page History
Content
| Table of Contents |
|---|
Phi-beta fit function
pyimgalgos/src/FiberAngles.py
y = funcy_l1_v1(x, phi_deg, bet_deg, DoR=392/913.27, sgnrt=-1.)
Two solutions of quadratic equation
- POS: y = -B + sqrt(B*B-C)
- NEG: y = -B - sqrt(B*B-C)
Legend:
- dot curves for negative beta
- black bold curve for beta=0
- solid curves for positive beta
Arc region
wide range of x:
narrow range of x:
Equatorial region
Problem with two roots is resolved for EQU region; sign of the root is selected the same as sign of parameter B;
- For |beta|>~48 solution does not exist for entire range of x
Data
exp=cxif5315:run=169
Fit in the processing script
Example of Here we discuss how to apply fit to data using absolute errors on peak position, get fit parameters with errors, estimate fit quality.
...
exp=cxif5315:run=169
Fit in the processing script
cxif5315/proc-cxif5315-r0169-peaks-from-file-v4.py, v5, v6
| Code Block | ||
|---|---|---|
| ||
from scipy.optimize import curve_fit
from scipy.stats import chi2
xn = np.array([pk.x/L for pk in sp.lst_equ_evt_peaks], dtype=np.double)
yn = np.array([pk.y/L for pk in sp.lst_equ_evt_peaks], dtype=np.double)
en = np.array([pk.csigma*sp.PIXEL_SIZE/L for pk in sp.lst_equ_evt_peaks], dtype=np.double)
#en = np.array([max(pk.rsigma,pk.csigma)*sp.PIXEL_SIZE/L for pk in sp.lst_equ_evt_peaks], dtype=np.double)
p0 = [-3.2,-16.0] # phi, beta angle central values
popt, pcov = curve_fit(funcy, xn, yn, p0, en, absolute_sigma=True)
sp.fib_chi2 = np.sum(((funcy(xn, *popt) - yn) / en)**2)
sp.fib_ndof = len(xn) - 1
sp.fib_prob = chi2.sf(sp.fib_chi2, sp.fib_ndof, loc=0, scale=1) |
...
Phi-beta fit results and quality
Distribution of parameters for peaks found by peak_finder_v2:
...
selected by Meng:
EQU region
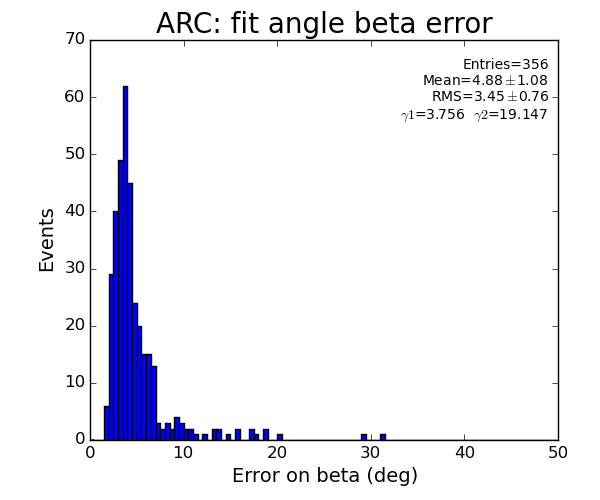
ARC region: 1-st solution for beta
ARC region: 2-nd solution for beta
For 2-peak events fit does not have any freedom - curve can exactly be fitted to 3 points (including origin); chi2=0 and probability is always 1.
For >2-peak events chi2 and probability show some distribution.
Peak list with phi-beta fit results
After phi-beta fit peaks are saved in the file with name like
...
| Note |
|---|
If someone want to load this file using |
References