Page History
| Table of Contents |
|---|
Motivation
Development of this application was stimulated by the discussion with Marcin Sikorski (meeting on
...
2012-08-30),
...
doing
...
xcs
...
experiments.
...
Users
...
need
...
in
...
real-time
...
algorithm
...
for
...
calculation
...
of
...
image
...
vs
...
time
...
auto-correlation
...
function
| Code Block |
|---|
} g2(tau) = <I(t)*I(t+tau)> / (<I(t)> * <I(t+tau)>), {code} where {{ |
where I(t)
...
is
...
an
...
image
...
intensity
...
at
...
time
...
t
...
,
...
and tau is a delay between two measurements.
Typical experimental condition can be described as follows:
- Run duration is about one hour at frequency up to 120 Hz that gives up to 10^5-10^6 images.
- Currently typical imaging devise is a Princeton camera with 1300x1340 pixels.
- Need to calculate
g2(tau)for each pixel, averaged over all possible image timestwith time differencetaubetween images. - A set of
taushould have about 30-100 points in log scale uniformly covering the run duration. - Use for example xcsi0112-r0015: 500 images with 8 sec delay between images.
Desired time for evaluation of the auto-correlation function should be comparable with run duration <1 hour. Currently this algorithm takes a few hours that can not be used for fast feedback in real time experiment.
Algorithm
Basic idea is (1) to split image vs time for small parts in image, (2) to process each part on separate computer node, (3) to merge results at the end of processing. It is clear that significant speedup (about T/N_nodes_) is achieved at the 2nd stage. These three stages are performed in separate C++ applications. Wrapping python script allows to submit job by a single command. It takes care about file and sub-process management in this job, as described below.
Code location
All modules for this application resides in the package ImgAlgos:
Module | Functionality |
|---|---|
ImgVsTimeSplitInFiles | splitter |
CorAna | base class with common methods |
CorAnaData | data processing for split files |
CorAnaInputParameters | provides storage for input parameters |
CorAnaMergeFiles | merging algorithm |
CorAnaProcResults | Example showing how to access results using C++ and produce a table for presentation |
CorAnaPars.py | singleton class for parameter storage in the wrapping file manager |
CorAnaSubmit.py | global methods for the file manager |
app/corana_submit | pythonic script which defines the sequence of procedures |
app/corana.cpp | main module for the part of image vs time correlation processing |
app/corana_merge.cpp | main module for merging |
app/corana_procres.cpp | main module for processing of results from correlator array |
data/psana-corana.cfg | psana configuration file for ImgVsTimeSplitInFiles |
data/PlotCorAnaResults.py | example of the python script which plots the resulting graphics |
Image splitting
Image splitting is implemented as a regular psana module ImgAlgos::ImgVsTimeSplitInFiles.
Command to run interactively on psana#### or submit in batch from pslogin## node:
| Code Block |
|---|
{{tau}} is a delay between two measurements. Typical experimental condition can be described as follows: * Run duration is about one hour at frequency up to 120 Hz that gives up to 10^5-10^6 images. * Currently typical imaging devise is a Princeton camera with 1300x1340 pixels. * Need to calculate {{g2(tau)}} for each pixel, averaged over all possible image times {{t}} with time difference {{tau}} between images. * A set of {{tau}} should have about 30-100 points in log scale uniformly covering the run duration. * Use for example xcsi0112-r0015: 500 images with 8 sec delay between images. Desired time for evaluation of the auto-correlation function should be comparable with run duration <1 hour. Currently this algorithm takes a few hours that can not be used for fast feedback in real time experiment. h1. More details 2012-09-10 meeting: In order to be useful this application should do correct math, accounts for image mask, discard bad events, noisy and "bright" pixels, apply normalization etc, and have a convenient GUI. Below is a list of requirements (marked as (?) ) with suggested solutions ( marked as (/) if exists or as (+) if needs to be implemented ). h3. Pedestals (?) "dark" run name should be provided by user and pedestals should be evaluated and applied for all runs until the "dark" run name has not changed. (/) For pedestals evaluation: use available {{ImgAlgos::ImgAverage}} psana module for "dark" run, which produces file with averaged over events pedestals (also produces the file with rms values). (/) For pedestals subtraction: use {{ImgAlgos::ImgCalib}} psana module right before evaluation of pedestals; the pedestals will be subtracted and corrected image will be retained in the event. h3. Low level threshold (?) Image pixel intensity physically can't be negative. Low amplitude noise should be suppressed by the threshold. The threshold amplitude should be provided by user (along with substituting amplitude). (+) Add this feature to the {{ImgAlgos::ImgCalib}} psana module, right after pedestal evaluation. h3. Image filtering (?) Usually users use different type of intensity monitor signals in order to retain/discard image for/from further processing. Discarded images should not contribute into the correlators evaluation. The spectra of the intensity monitors should be available for browsing. User should be able to select the intensity monitor(s) from the list and set low and high thresholds. (+) The filtering module may be implemented in psana. Based on selected intensity monitor(s) and thresholds it will decide to retain or discard event and accumulate spectral histograms. The histograms will be saved in file at the end of run. (+) Control GUI should be able to browse the intensity monitor histograms and set the thresholds. h3. Selection of intensity monitors (?) It would be nice to have an algorithm like in XTC explorer (+) Possible options: * run application as a plug-in for XTC Explorer, * pyana modeule performing similar to XTC Explorer algorithm, * stand-alone C\+\+ module reading XTC datagrams, * hardwired list of intensity monitors. h3. Dynamic mask (?) Imaging camera may have permanently hot pixels or some pixels may be saturated during the run. User need to set a threshold on high intensity. If the pixel amplitude crosses this threshold at least once during the run, then this pixel should be excluded from further analysis. The same is valid for hot pixels, which shows non-zero intensity in large fraction of events. (+) This can be done in psana module, which works before event selection algorithm. Two files of image size may be produced 1) for satturated and 2) for hot pixels. h3. Static mask (?) The beam-stopper region and some areas with fringes should be masked. It could be useful to have a graphical editor for mask. (+) See section for GUI h3. Graphical editor for selected regions (?) Sometimes it is useful to restrict good region of the image (+) See section for GUI h3. Center of the image (?) User should have an option to set a center of the rings for histograms. (+) This could be done at the final stage, when all correlatirs are available. (+) See section for GUI h3. Correct normalization of g2 (?) Evaluation of {{g2}} for image regions is not that simple as presented by the fomulae for a single pixel: {code} g2(tau) = <I(t)*I(t+tau)> / (<I(t)> * <I(t+tau)>), {code} In order to get physically meaningful results for g2, the correlators <I(t)> and <I(t+tau)> should be averaged in the fine rings around center with number of bins N2, which is order of 100, with dR down to 1-2 pixels. Then the <I(t)*I(t+tau)> (?) correlator should be averaged over bold rings intended for G2 evaluation. The number of these rings N1 should be order of 10. The N2 and N1 should be defined by user. It might be useful to define the histogram region by the sector in the user-defined angular range. (+) See section for GUI h3. GUI (?) In order to get an easy interface to all subprocesses, it seems useful to have a GUI with configuration of everything through the GUI. (!) Well, presumably users will want different specific features in their analyses which can not be foreseen in implementation of the GUI. It is pretty unlikely that everything in analysis can be done clicking on buttuns in GUI. Then, it could be nice if user understand what he is doing step by step and have a monitoring at the end of each stage. We are doing science, not a standard pre-defined things... Most generic way to process data is to have a separate procedures with command line interface. (+) Anyway, the browser/presenter of data stored in the files after pre-processing could be provided for a set of common specific plots. All features listed in previous sections, such as static and dynamic mask, restriction of the region(s) of interest, selection of the image center, the binning scheme etc., can be done in the browser at the final stage of the analysis. ---- ---- ---- ---- ---- ---- {HTMLcomment:hidden} Here is my comment {HTMLcomment} h1. Algorithm Basic idea is (1) to split image vs time for small parts in image, (2) to process each part on separate computer node, (3) to merge results at the end of processing. It is clear that significant speedup (about T/N_nodes_) is achieved at the 2nd stage. These three stages are performed in separate C+\+ applications. Wrapping python script allows to submit job by a single command. It takes care about file and sub-process management in this job, as described below. h3. Code location All modules for this application resides in the [package ImgAlgos|PCDS:Psana Module Catalog#Package ImgAlgos]: || Module || Functionality || | ImgVsTimeSplitInFiles | splitter | | CorAna | base class with common methods | | CorAnaData | data processing for split files | | CorAnaInputParameters | provides storage for input parameters | | CorAnaMergeFiles | merging algorithm | | CorAnaProcResults | Example showing how to access results using C+\+ and produce a table for presentation | | CorAnaPars.py | singleton class for parameter storage in the wrapping file manager | | CorAnaSubmit.py | global methods for the file manager | | app/corana_submit | pythonic script which defines the sequence of procedures | | app/corana.cpp | main module for the part of image vs time correlation processing | | app/corana_merge.cpp | main module for merging | | app/corana_procres.cpp | main module for processing of results from correlator array | | data/psana-corana.cfg | psana configuration file for ImgVsTimeSplitInFiles | | data/PlotCorAnaResults.py | example of the python script which plots the resulting graphics | h3. Image splitting Image splitting is implemented as a regular psana module [ImgAlgos::ImgVsTimeSplitInFiles|PCDS:Psana Module Catalog#Module ImgAlgos::ImgVsTimeSplitInFiles]. Command to run interactively on {{psana####}} or submit in batch from {{pslogin##}} node: {code} psana -c <config-file> <xtc-file-list> bsub -q psfehq -o log-file 'psana -c <config-file> <xtc-file-list>' {code} |
For
...
example:
| Code Block |
|---|
} psana -c ImgAlgos/data/psana-corana.cfg /reg/d/psdm/XCS/xcsi0112/xtc/e167-r0015-* {code}* |
where
...
ImgAlgos/data/psana-corana.cfg
...
is
...
an
...
example
...
of
...
the
...
configuration
...
script
...
for
...
psana
...
and
...
/reg/d/psdm/XCS/xcsi0112/xtc/e167-r0015-
...
*
...
are
...
the
...
input
...
xtc
...
files
...
for
...
particular
...
run.
| Note |
|---|
A couple of limitations due to LCLS policy: |
Produces the files:
| Code Block |
|---|
{note} A couple of limitations due to LCLS policy: Interactive job can be run on {{psana####}} computer, but the batch queues are not seen from {{psana####}} nodes... Batch job can be submitted from {{pslogin##}} computer, but data are not seen directly from {{pslogin##}} nodes... {note} Produces the files: {code} cor-ana-r0015-b0000.bin - file with a part of image vs time cor-ana-r0015-b0001.bin cor-ana-r0015-b0002.bin cor-ana-r0015-b0003.bin cor-ana-r0015-b0004.bin cor-ana-r0015-b0005.bin cor-ana-r0015-b0006.bin cor-ana-r0015-b0007.bin cor-ana-r0015-time.txt - list of time-records for all events in processed run. cor-ana-r0015-time-ind.txt - list of time-records for all events in processed run with time index. cor-ana-r0015-med.txt - file with metadata. In particular it has the original image size, number of image parts for splitting, number of images in run, etc. {code} |
Algorithms:
...
- The
...
- <int16_t>
...
- image
...
- data
...
- array
...
- is
...
- split
...
- for
...
- ordered
...
- number
...
- of
...
- equal
...
- parts
...
- (by
...
- the
...
- parameters
...
nfiles_out
...
- in
...
- psana-corana.cfg
...
- file)
...
- and
...
- each
...
- part
...
- is
...
- saved
...
- in
...
- the
...
- output
...
cor-ana-r0015-b####.bin
...
- file
...
- sequentially
...
- for
...
- all
...
- selected
...
- events.
...
- The
...
- appropriate
...
- time
...
- record
...
- for
...
- selected
...
- event
...
- is
...
- saved
...
- in
...
- the
...
- file
...
cor-ana-r0015-time.txt
...
- .
...
- At
...
- the
...
- end
...
- of
...
- the
...
- splitting
...
- procedure:
...
- the
...
- average
...
- time
...
- difference
...
- and
...
- its
...
- rms
...
- between
...
- sequential
...
- events
...
- is
...
- evaluated
...
- for
...
- all
...
- recorded
...
- time
...
- records.
...
- The
...
- file
...
cor-ana-r0015-time.txt
...
- is
...
- re-processed
...
- and
...
- for
...
- each
...
- record
...
- the
...
- time
...
- index
...
- is
...
- evaluated
...
- as
...
- unsigned
...
- value
...
- of
...
Code Block
...
<time-index> = (<event-time> + 0.5 <average-time-between-events>) / <average-time-between-events>
...
- Event record with time index is saved in the file
cor-ana-r0015-time-ind.txt
...
- All
...
- metadata
...
- parameters
...
- which
...
- are
...
- required
...
- for
...
- further
...
- processing,
...
- such
...
- as
...
- input
...
- parameters,
...
- image
...
- size,
...
<average-time-between-events
...
- ,
...
- maximal
...
- value
...
- of
...
- the
...
- time
...
- index
...
- etc.,
...
- are
...
- saved
...
- in
...
- file
...
cor-ana-r0015-med.txt
...
- .
| Note |
|---|
This approach allows to apply the modest event selection algorithms in |
Time correlation processing
ImgAlgos/app/corana application
Command to run interactively on psana#### or submit in batch from pslogin## node:
| Code Block |
|---|
{note} This approach allows to apply the modest event selection algorithms in {{psana}} pre-processing stage. But, it still based on uniform time indexing... Q: Is it really good assumption for this kind of experiments? {note} h3. Time correlation processing {{ImgAlgos/app/corana}} application Command to run interactively on {{psana####}} or submit in batch from {{pslogin##}} node: {code} corana -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h] bsub -q psfehq -o log-file 'corana -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h]' {code} |
For
...
example
...
the
...
interactive
...
and
...
batch
...
mode
...
commands:
| Code Block |
|---|
} corana -f cor-ana-r0015-b0001.bin -t my-tau.txt bsub -q psfehq -o log-file 'corana -f cor-ana-r0015-b0000.bin' {code} |
Produce
...
files:
| Code Block |
|---|
} cor-ana-r0015-tau.txt - string of {{tau}} values for which the auto-correlation function is evaluated cor-ana-r0015-b0000-result.bin - auto-correlators for the part of the image for all {{tau}} values cor-ana-r0015-b0001-result.bin cor-ana-r0015-b0002-result.bin cor-ana-r0015-b0003-result.bin cor-ana-r0015-b0004-result.bin cor-ana-r0015-b0005-result.bin cor-ana-r0015-b0006-result.bin cor-ana-r0015-b0007-result.bin {code} h3. Merging results {{ |
Merging results
ImgAlgos/app/corana_merge
...
application
...
Command
...
to
...
run
...
interactively
...
on
...
psana####
...
or
...
submit
...
in
...
batch
...
from
...
pslogin##
...
node:
| Code Block |
|---|
} corana_merge -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h] bsub -q psfehq -o log-file 'corana_merge -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h]' {code} |
For
...
example:
| Code Block |
|---|
} corana_merge -f cor-ana-r0015-b0001-result.bin -t my-tau.txt {code} |
This
...
procedure
...
produces
...
file:
| Code Block |
|---|
} cor-ana-r0015-image-result.bin {code} h3. Example of how to get and process results {{ |
Example of how to get and process results
ImgAlgos/app/corana_procres
...
Command
...
to
...
run
...
interactively
...
on
...
psana####
...
or
...
submit
...
in
...
batch
...
from
...
pslogin##
...
node:
| Code Block |
|---|
} corana_procres -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h] bsub -q psfehq -o log-file 'corana_procres -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h]' {code} |
Basically
...
it
...
reads
...
files
...
with
...
results
...
and
...
produces
...
the
...
histogram-like
...
table
...
*-hist.txt
...
.
Automatic processing
ImgAlgos/app/corana_submit
...
-
...
is
...
a
...
wrapping
...
script
...
which
...
allows
...
to
...
run
...
all
...
of
...
above
...
procedures
...
by
...
a
...
single
...
command
...
from
...
pslogin##
...
node
...
and
...
it
...
keeps
...
eye
...
on
...
processing
...
of
...
jobs
...
in
...
batch
...
and
...
doing
...
the
...
file
...
management.
...
Command
...
to
...
start:
| Code Block |
|---|
} corana_submit [-c <config-file>] [-t <fname-tau>] [-x] <xtc-file-list> {code} |
For
...
example:
| Code Block |
|---|
} corana_submit -c ImgAlgos/data/psana-corana.cfg -t my-tau.txt /reg/d/psdm/XCS/xcsi0112/xtc/e167-r0015-s00-c00.xtc {code} |
This
...
script
...
sequentially
...
performs
...
operations
...
for
...
single
...
run
...
as
...
follows:
...
- Initialize
...
- all
...
- parameters
...
- Run
...
- psana
...
- to
...
- split
...
- image
...
- for
...
- files
...
- Check
...
- that
...
- all
...
- split
...
- files
...
- are
...
- produced
...
- Submit
...
- job
...
- for
...
- time-correlation
...
- processing
...
- Check
...
- that
...
- all
...
- processed
...
- files
...
- are
...
- produced
...
- Submit
...
- job
...
- for
...
- merging
...
- Check
...
- that
...
- merged
...
- file
...
- is
...
- produced
...
- Submit
...
- job
...
- for
...
- test
...
- processing
...
- of
...
- the
...
- file
...
- with
...
- results
...
- List
...
- all
...
- created
...
- files
...
- Clean-up
...
- files
...
- in
...
- the
...
- work
...
- directory
...
- List
...
- of
...
- preserved
...
- files
...
Note
...
The
...
next
...
to
...
last
...
procedure
...
deletes
...
all
...
intermediate
...
split
...
-
...
and
...
log
...
-
...
files.
...
In
...
debugging
...
mode
...
this
...
procedure
...
may
...
be
...
turned
...
off.
...
Manual sequential processing
In case of manual processing of all scripts, commands need to be issued in a right order. Commands corana, corana_merge, and corana_procres should have the same list of parameters. This is important, because all file names for these procedures are generated by the same base class ImgAlgos/src/CorAna.cpp
...
Right
...
sequence
...
of
...
commands
...
to
...
run
...
interactively
...
on psana####
| Code Block |
|---|
{{psana####}} {code} psana -c <config-file> <xtc-file-list> corana -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h] corana_merge -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h] corana_procres -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h] {code} |
or
...
submit
...
in
...
batch
...
from
...
pslogin##
...
node:
| Code Block |
|---|
} bsub -q psfehq -o log-file 'psana -c <config-file> <xtc-file-list>' bsub -q psfehq -o log-file 'corana -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h]' bsub -q psfehq -o log-file 'corana_merge -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h]' bsub -q psfehq -o log-file 'corana_procres -f <fname-data> [-t <fname-tau>] [-l <logfile>] [-h]' {code} The {{corana}} batch jobs can be submitted and run on separate butch nodes in parallel. All other procedures can be submitted when previous is successfully finished and all necessary files are produced. The {{corana_procres}} command is optional and is currently used for test purpose only. But, it may be replaced by real analysis code. h3. File formats * File with split-image data for selected events {{ |
The corana batch jobs can be submitted and run on separate butch nodes in parallel. All other procedures can be submitted when previous is successfully finished and all necessary files are produced.
The corana_procres command is optional and is currently used for test purpose only. But, it may be replaced by real analysis code.
File formats
- File with split-image data for selected events
cor-ana-r0015-b000N.bin
...
- :
...
Currently
...
- this
...
- file
...
- contains
...
<uint16_t>
...
- amplitude
...
- for
...
- each
...
- pixel
...
- in
...
- binary
...
- format
...
- for:
...
Code Block
...
<data-for-img-partN-of-img1> <data-for-img-partN-of-img2> ... <data-for-img-partN-of-imgLast>
...
- File with metadata parameters
cor-ana-r0015-med.txt
...
- :
...
Code Block
...
IMAGE_ROWS 1300 IMAGE_COLS 1340 IMAGE_SIZE 1742000 NUMBER_OF_FILES 8 BLOCK_SIZE 217750 REST_SIZE 0 NUMBER_OF_IMGS 500 FILE_TYPE bin DATA_TYPE uint16_t TIME_SEC_AVE 8.088413 TIME_SEC_RMS 0.063639 TIME_INDEX_MAX 499
...
- File with image time records
cor-ana-r0015-time.txt
...
- :
...
Code Block
...
1 0.000000 0.000000 20120616-080236.671607864 5366 0 2 8.026429 8.026429 20120616-080244.698036743 8255 1 3 16.144788 8.118359 20120616-080252.816395836 11177 2 4 24.154835 8.010048 20120616-080300.826443448 14060 3 ...
...
where
...
- each
...
- record
...
- has:
...
Code Block
...
<image-in-file#> <t(sec)-from-the-1st-event> <dt(sec)> <time-stamp> <fiducials> <event#-since-configure>
...
- File with image time records and evaluated time index
cor-ana-r0015-time-ind.txt
...
- :
...
Code Block
...
1 0.000000 0.000000 20120616-080236.671607864 5366 0 0 2 8.026429 8.026429 20120616-080244.698036743 8255 1 1 3 16.144788 8.118359 20120616-080252.816395836 11177 2 2 4 24.154835 8.010048 20120616-080300.826443448 14060 3 3 5 32.281937 8.127102 20120616-080308.953545010 16985 4 4 ...
...
where
...
- each
...
- record
...
- has:
...
Code Block
...
<image-in-file#> <t(sec)-from-the-1st-event> <dt(sec)> <time-stamp> <fiducials> <event#-since-configure> <time-index-starting-from-0>
...
- File with split-image
...
- correlators
...
- for
...
- each
...
- value
...
- of
...
tau
...
cor-ana-r0015-b000N-result.bin
...
- :
...
Currently
...
- it
...
- saves
...
<float>
...
- correlator
...
- for
...
- each
...
- pixel
...
- in
...
- binary
...
- format
...
- for:
...
Code Block
...
<corr-for-img-partN-of-tau1> <corr-for-img-partN-of-tau2> ... <corr-for-img-partN-of-tauLast>
...
my-tau.txt
...
- :
...
Code Block
...
1 3 5 7 9 10 12 14 16 18 20 24 28 30 32 36 40 ... 160 180 200 240 280 300 320 360 400
...
contains
...
- the
...
tau
...
- values
...
- presented
...
- in
...
- terms
...
- of
...
- number
...
- of
...
- ordered
...
- images
...
- in
...
- the
...
- file.
Quick start guide
We assume that everything is set up to work on LCLS analysis farm, otherwise see Computing (including Analysis) and Account Setup.
How to run this procedure
If the version of the package ImgAlgos is available as a current software release, then you may run the script command(s) directly, for example:
| Code Block |
|---|
h1. Quick start guide We assume that everything is set up to work on LCLS analysis farm, otherwise see [PCDS:Computing] and [Account Setup|PCDS:Analysis Workbook. Account Setup]. h3. How to run this procedure If the version of the [package ImgAlgos|PCDS:Psana Module Catalog#Package ImgAlgos] is available as a current software release, then you may run the script command(s) directly, for example: {code} cd <your-favorite-directory> mkdir work_corana sit_setup corana_submit [-c <config-file>] [-t <fname-tau>] [-x] <xtc-file-list> {code} {note} If the code in the [package ImgAlgos|PCDS:Psana Module Catalog#Package ImgAlgos] has been recently changed and the updated release is not yet available, then one need to create the local release directory, get the latest/HEAD version of the package, and compile the code as shown below: {note} {code} |
| Note |
|---|
If the code in the package ImgAlgos has been recently changed and the updated release is not yet available, then one need to create the local release directory, get the latest/HEAD version of the package, and compile the code as shown below: |
| Code Block |
|---|
cd <your-favorite-directory>
newrel ana-current myReleaseDirectory
cd myReleaseDirectory
sit_setup
addpkg ImgAlgos HEAD
scons
{code}
h3. Where to find results
The procedure will produce a bunch of files in the {{work_corana}} directory. If everything is OK, then all spit - and log\- files will be removed at the end of automatic {{corana_submit}} procedure. The most important files are preserved for further analysis:
|| File name tail || Format || Content ||
| \ |
Where to find results
The procedure will produce a bunch of files in the work_corana directory. If everything is OK, then all spit - and log- files will be removed at the end of automatic corana_submit procedure. The most important files are preserved for further analysis:
File name tail | Format | Content |
|---|---|---|
*-image-result.bin |
...
binary |
...
for |
...
<float> |
...
correlators |
...
for |
...
all |
...
image |
...
pixels |
...
for |
...
all |
...
tau |
...
values |
*-time-ind.txt |
...
text | time records for all selected events/images | |
*-tau.txt |
...
text | the list of tau intervals | |
*-med.txt |
...
text | meta data parameters | |
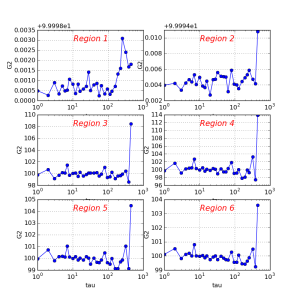
*-hist.txt | text | Histogram array with correlators averaged for ring regions of the image for all |
How to look at results
It is assumed that all files listed in previous section may be used for further analysis, depending on particular goals. The optional script corana_procres is designed as an example of how to access data from C++ code. Class CorAnaProcResults produces the file *-hist.txt
A simple python script shows how to plot this file:
| Code Block |
|---|
| text | Histogram array with correlators averaged for ring regions of the image for all {{tau}} values, shown in the first column | h3. How to look at results It is assumed that all files listed in previous section may be used for further analysis, depending on particular goals. The optional script {{corana_procres}} is designed as an example of how to access data from C+\+ code. Class {{CorAnaProcResults}} produces the file {{\*-hist.txt}} A simple python script shows how to plot this file: {code} ./ImgAlgos/data/PlotCorAnaResults.py work_corana/cor-ana-r0015-hist.txt {code} !image.png|thumbnail,border=1! {note} Another option is to use python script for direct processing of the resulting files. This is not elaborated yet. Q: What kind of further processing is desired and what tools are going to be used? {note} |
| Note |
|---|
Another option is to use python script for direct processing of the resulting files. |