"Data analysis == Piece of cake"
Sections in this tutorial
It is always a good idea for the people doing analysis to be able to look at their detector images and probe intensity values. Given that a typical LCLS experiment has millions of snapshots to choose from, it is also critical that you can quickly select images of interest and set regions of interest using masks. By the end of this tutorial, you will be able to browse images, jump to images of interest, generate masks, find peaks in your images and index crystal diffraction patterns.
Citation for psocake (and other psana-based programs):
@article{Damiani:zw5004,
author = "Damiani, D. and Dubrovin, M. and Gaponenko, I. and Kroeger, W. and Lane, T. J. and Mitra, A.
and O'Grady, C. P. and Salnikov, A. and Sanchez-Gonzalez, A. and Schneider, D. and Yoon, C. H.",
title = "{Linac Coherent Light Source data analysis using {it psana}}",
journal = "Journal of Applied Crystallography",
year = "2016", volume = "49", number = "2", pages = "672--679", month = "Apr",
doi = {10.1107/S1600576716004349}, url = {http://dx.doi.org/10.1107/S1600576716004349}, } |
Type "psocake" on your terminal to open up the GUI. For crystallography, we will need to open it in sfx mode (-m):
$ sit_setup nightly-20160717 # (Optional) To get the bleeding edge version, use the nightly build $ psocake -m sfx |
1) There are four parameters required to uniquely identify an image at LCLS. Type the experiment name, run number, detector name, and event number in the Experiment Parameters panel.
For this tutorial, we will look at experiment cxitut13, run 10, detector DscCsPad, event 11.
#######################################
# Available area detectors:
# ('CxiDs1.0:Cspad.0', 'DscCsPad', '')
####################################### |
2) You can specify the experiment parameters as command line arguments in psocake using the psana-style experiment run string:
$ psocake exp=cxitut13:run=10 -d DscCsPad -n 11 -m sfx |
3) You can also use the -e and -r arguments for the experiment and the run number:
$ psocake -e cxitut13 -r 10 -d DscCsPad -n 11 -m sfx |
To check psocake version:
$ psocake --version |
Don’t worry if you don’t remember these arguments. For argument options, use --help:
$ psocake --help |
In this example, let's mask out pixels that should not be used for analysis, mask out the jet streak at the centre of the detector, and mask out the water ring.
Note: the Image Panel must be in the default "greyscale" colormap for the mask colors to display properly.
1) On the mask panel, click on "Use psana mask". This will mask out the following pixels that should not be used for analysis; calib, status, edge, central, unbonded pixels, unbonded pixel neighbor pixels. These pixels are shown as green on the image panel.

2) On the mask panel, click on "Use streak mask". This will mask out strong intensities originating from the edges of the central asics.
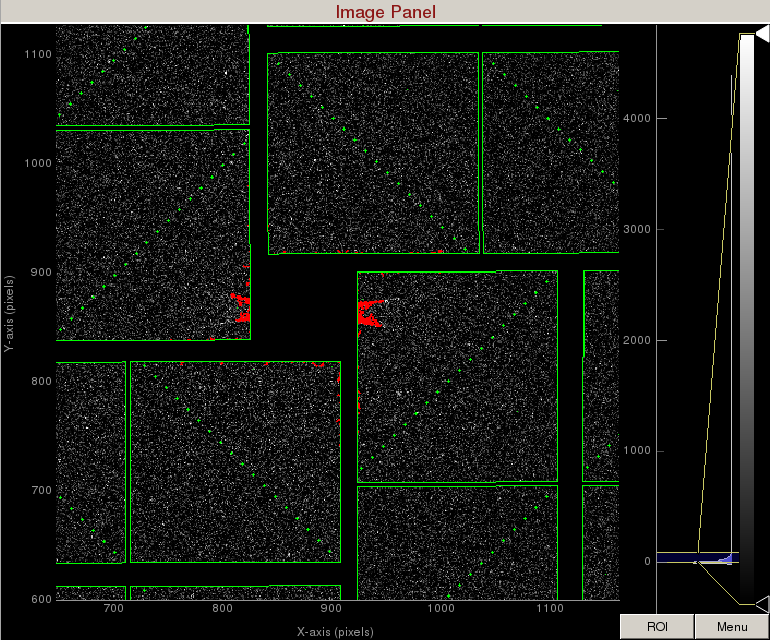
3) To make a donut mask over the water ring, click on "Use user-defined mask". This will bring up a cyan circle and cyan square mask generator.
Select "Toggle" in Masking mode. Move the cyan circle to the centre of the detector by dragging the circle. Resize the cyan circle by dragging the diamond on the perimeter. Once you are happy with the position, click "mask circular ROI" button on the mask panel.
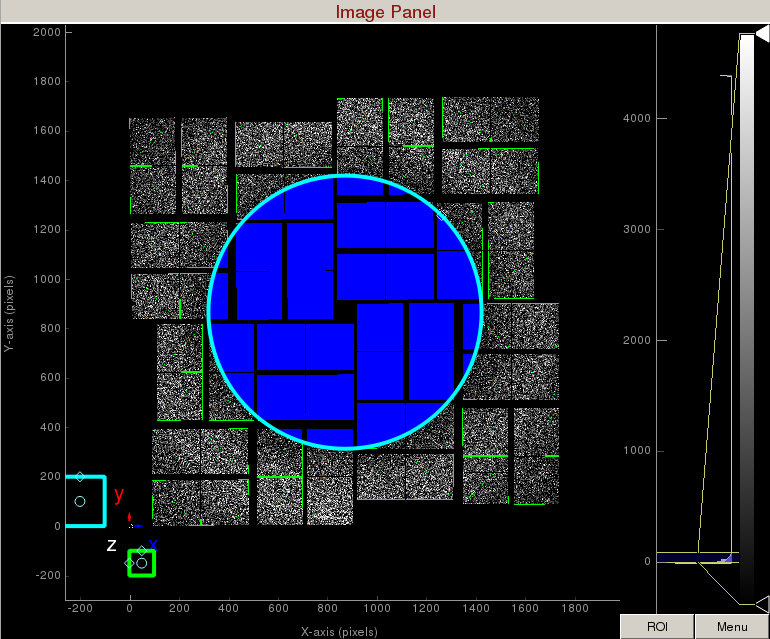
Increase the cyan circle again by dragging the diamond on the perimeter. Click "mask circular ROI" button on the mask panel. Because we are in the "toggle" mode, the previous mask gets toggled and disappears. The area that does not overlap with the previous mask get masked out.
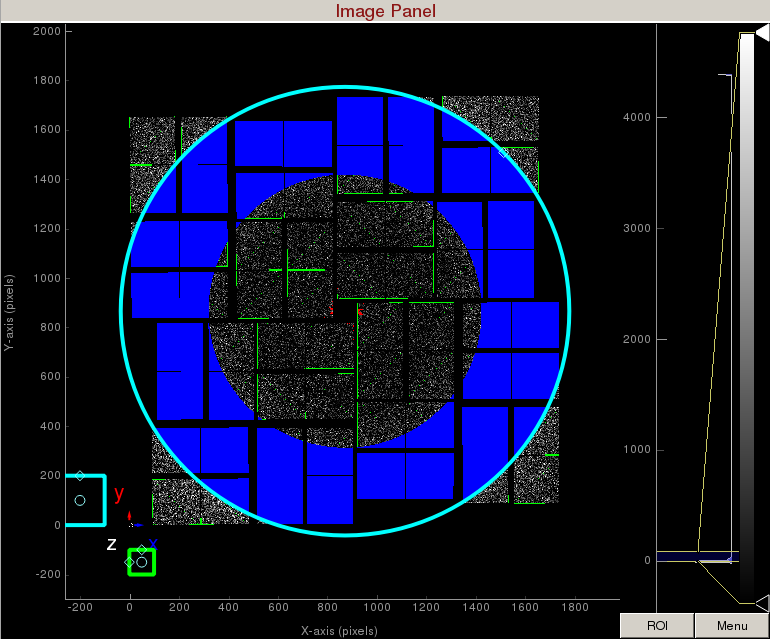
To save the user-defined mask, click on "Save user-defined mask" on the mask panel which will save the mask in the scratch folder. For this example, the mask will be saved here:
/reg/d/psdm/cxi/cxitut13/scratch/<username>/psocake/r0010/mask.npy (3D ndarray)
/reg/d/psdm/cxi/cxitut13/scratch/<username>/psocake/r0010/mask.txt (2D text)
You can load the user-defined mask using the "Load user-defined mask" button and selecting mask.npy.
Hover the mouse pointer over the Bragg peaks to study the intensities. The sum of the Bragg peak pixels are above 400 ADUs. Set the following values:
The output directory on the peak finder panel should already be automatically set to: /reg/d/psdm/cxi/cxitut13/scratch/<username>/psocake
In the small data panel, type the CXIDB filename:
First things first, crystal indexing requires an accurate detector geometry. Latest CXI geometry files can be found here: Geometry history
Detector panels can manually adjusted using: calibman
If you are on a psana machine, you can run CrystFEL programs by setting up your environment:
source /reg/g/cfel/crystfel/crystfel-dev/setup-sh
CFDEPDIR=/reg/g/cfel/dependencies
export PATH=${CFDEPDIR}/bin:$PATH |
Let's try to index the diffraction pattern at event 11.
In the indexing panel, tick "Indexing on". Indexed peaks will be marked with magenta circles. These triple rings represent the integration radius.
Hopefully, you have indexed this diffraction pattern. Notice that the unitcell parameters are a bit off compared to what is expected. Let's load a CrystFEL unitcell file to help the indexer along.
CrystFEL unit cell file version 1.0 lattice_type = tetragonal centering = P unique_axis = c a = 77.05 A b = 77.05 A c = 37.21 A al = 90 deg be = 90 deg ga = 90 deg |
If you would like to save the detector images in the .cxi file, turn on "Keep CXI images". Only set this to true, if you anticipate that you will want to reindex this run. Otherwise, it's just a total waste of your precious disk space.
As with peak finding, you can launch indexing jobs on multiple runs by specifying runs in the Run(s) field.
Indexing will take some time to complete. If successful, you should see a stream file in: /reg/d/psdm/cxi/cxitut13/scratch/<username>/psocake/r0010/cxitut13_10.stream
In the small data panel, type the CXIDB filename:
Please send bug reports/comments:
yoon82@slac.stanford.edu